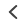Structured illumination is a computational imaging system that involves the projection of a known pattern of light onto a sample; the way the pattern deforms when the projected light strikes different components in the sample allows imaging systems to calculate additional information about the sample. This technique is used in several technologies, including a popular forensic imaging device, the 3D scanner. In three-dimensional structured illumination microscopy (3D SIM), a fluorescently labeled microscopic sample is excited with structured light, providing information outside the diffraction limit. Computational interpretation of this information yields a super-resolution image with double the resolution of standard light microscopy. Though these gains are modest among super-resolution methods, this technique offers advantages in improved optical sectioning, lower required illumination dose and acquisition speed, and has provided numerous biological insights. One limitation of 3D SIM is its axial resolution, which is normally limited to ∼300-nm, versus ∼120-nm lateral resolution, leading to distortion of fine features along the axial dimension.
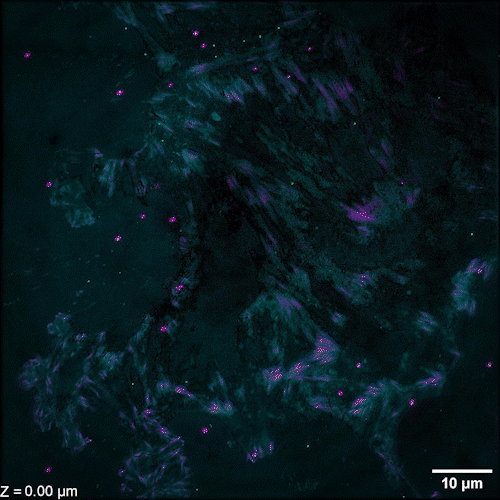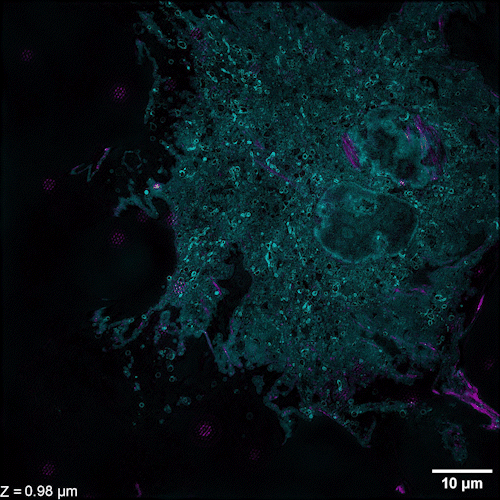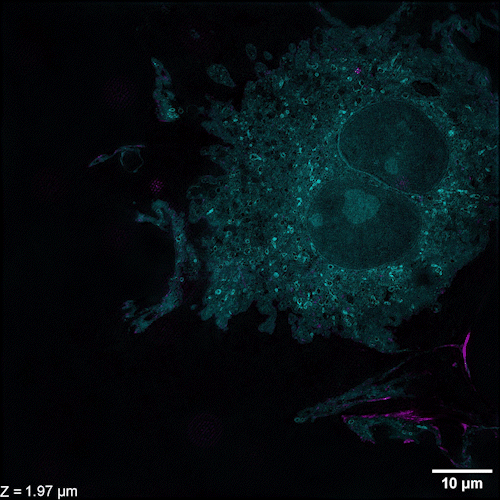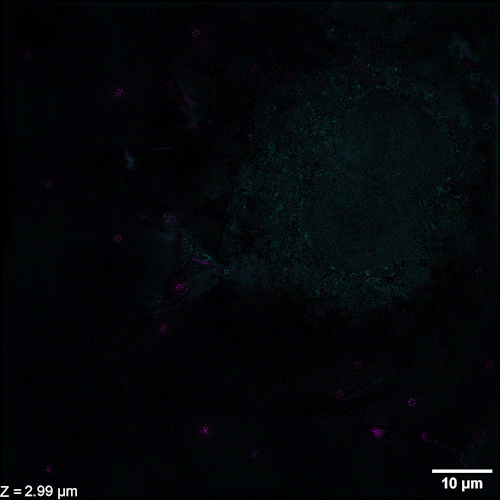
Li et al. describe methods of overcoming this issue with 3D SIM axial resolution in their recent Nature Biotechnology article, which includes several striking images. They present two discrete, complementary methods to improve axial resolution, which require little or no alteration to existing optical systems. They first showed that placing a mirror on the opposite side of the sample and back-reflecting the central beam enables four-beam interference offering near-isotropic imaging. They also developed a multi-step deep learning pipeline that produced reconstructions with isotropic spatial resolution. They then use a panel of dyes, including Biotium’s CellBrite® Fix 488 and CellBrite® Fix 555 fixable membrane stains, to demonstrate the potential of their novel methods by imaging a variety of prokaryotic and eukaryotic cellular samples and delineating the nanoscale distribution of subcellular elements. In bacteria, they were able to resolve the structure of membranes, cell division proteins, and core components of the spore coat (Figure 1 a-d; Video 1). In eukaryotic cells, they achieved nanoscale resolution of membrane-encased actin filaments and pores that traversed thin membrane extensions, discrete cytoskeletal elements, and caveolar coat proteins. They also performed time-lapse imaging of organellar and cytoskeletal dynamics (Figure 1 h-n; Video 2). Together these results demonstrate the utility of these two approaches for improving 3D SIM. Such advances are important, as SIM microscopy offers a potential replacement for electron microscopy for some medical diagnoses, including those related to kidney disorders, kidney cancer, and blood diseases.
Videos

Learn more about Biotium’s broad selection of membrane dyes and high-performance CF® Dyes with multi-color flexibility, along with other reagents for immunofluorescence microscopy. We also offer several product lines that have been validated for STORM and other super-resolution microscopy techniques.
Full Citation
Li, X., Wu, Y., Su, Y., Rey-Suarez, I., Matthaeus, C., Updegrove, T. B., … & Shroff, H. (2023). Three-dimensional structured illumination microscopy with enhanced axial resolution. Nature Biotechnology, 1-13. https://doi.org/10.1038/s41587-022-01651-1
Other References
Pullman, J. M., Nylk, J., Campbell, E. C., Gunn-Moore, F. J., Prystowsky, M. B., & Dholakia, K. (2016). Visualization of podocyte substructure with structured illumination microscopy (SIM): a new approach to nephrotic disease. Biomedical Optics Express, 7(2), 302-311. doi:10.1364/BOE.7.000302
Liu, J., Wang, M., Tulman, D., Mandava, S. H., Elfer, K. N., Gabrielson, A., … & Lee, B. R. (2016). Nondestructive diagnosis of kidney cancer on 18-gauge core needle renal biopsy using dual-color fluorescence structured illumination microscopy. Urology, 98, 195-199. doi:10.1016/j.urology.2016.08.036
Westmoreland, D., Shaw, M., Grimes, W., Metcalf, D. J., Burden, J. J., Gomez, K., … & Cutler, D. F. (2016). Super‐resolution microscopy as a potential approach to diagnosis of platelet granule disorders. Journal of Thrombosis and Haemostasis, 14(4), 839-849. doi:10.1111/jth.13269