PMAxx™ Dye, 20 mM in H2O
Viability PCR dye, a new and improved version of the popular viability dye propidium monoazide (PMA).
Please fill in the inquiry form and we will contact you shortly.
Wishlist updated! View wishlist
Product Description
PMAxx™ is a photoreactive DNA-binding dye for viability PCR. It was designed by Biotium scientists to be a superior alternative to the popular viability dye propidium monoazide (PMA).
- Selectively detect viable cells using qPCR
- Better live/dead discrimination of bacteria than PMA
- Dead cell specific dye, binds to DNA
- Covalently attaches to DNA after photoactivation
- 20 mM in water, 100 uL volume
- λAbs = 464 nm (before photolysis); λAbs/λEm= ~510/~610 nm (with DNA/RNA, after photolysis)
To learn more about the advantages of determining microbial or cell viability using viability PCR, visit the Viability PCR Technology Page.
About PMAxx™
PMAxx™ dye is a DNA modifier used for viability PCR, invented by scientists at Biotium. PMAxx™ is a new and improved version of our popular viability dye PMA (propidium monoazide). Like PMA, PMAxx™ is a photo-reactive dye that binds to DNA with high affinity. Upon photolysis with visible light, PMAxx™ dye becomes covalently attached to DNA. This modified DNA cannot be amplified by PCR. PMAxx™ dye is designed to be cell membrane-impermeant. Thus, in a population of live and dead cells, only dead cells are susceptible to DNA modification due to compromised cell membranes. This unique feature makes PMAxx™ highly useful in selective detection of live bacteria by qPCR.
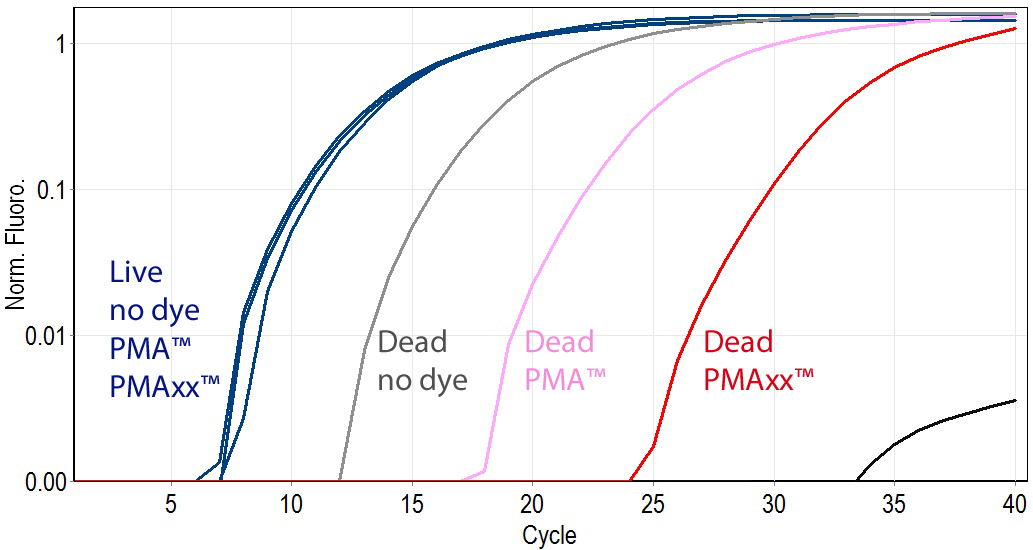
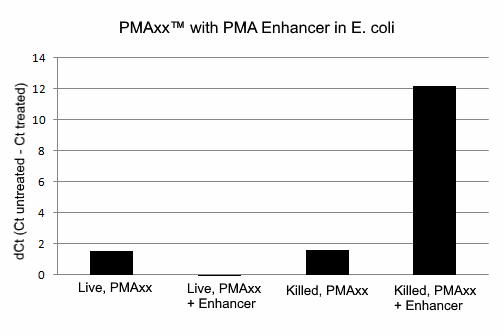
PMAxx™ was designed by Biotium scientists to be a superior alternative to PMA, and is compatible with all existing PMA protocols, but with superior activity. While PMA is generally effective at differentiating between live and dead bacteria by qPCR, it does not completely eliminate PCR products from dead cell DNA. This could potentially give false positive results. Biotium’s new dye PMAxx™ is much more effective at eliminating PCR amplification of dead cell DNA, and therefore provides the best discrimination between live and dead bacteria.
References
Download list of curated PMA and PMAxx™ References and a list of PMA and PMAxx™ Validated Bacterial Strains.