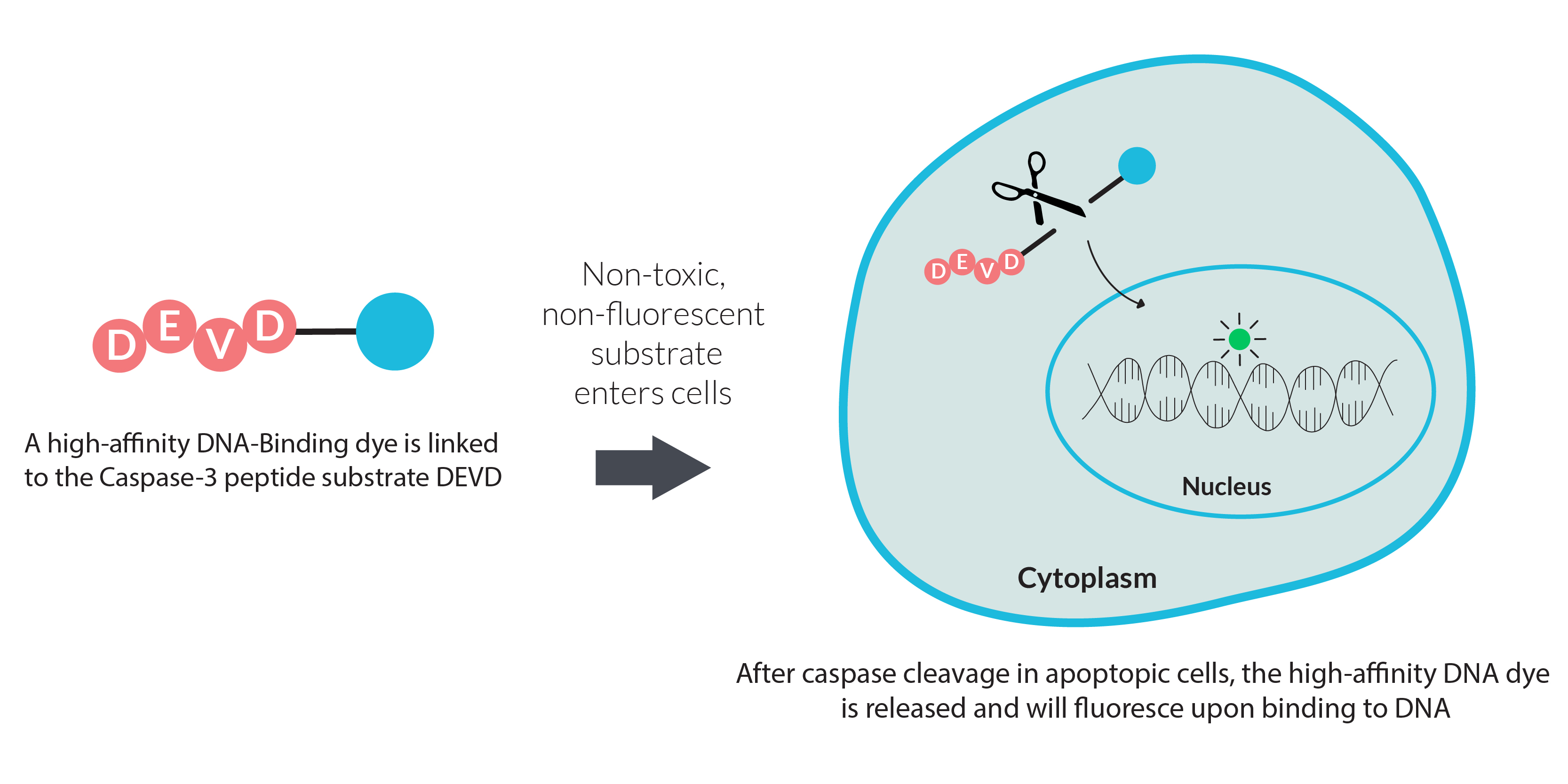
How It Works
Features
- Dual detection of caspase activity and nuclear morphology
- Rapid, no-wash, endpoint or real-time assays
- Non-toxic, allowing for multi-day experiments
- For flow cytometry, microscopy, or live cell imaging systems
- Available as stand-alone substrates or in kits paired with other apoptosis or necrosis probes

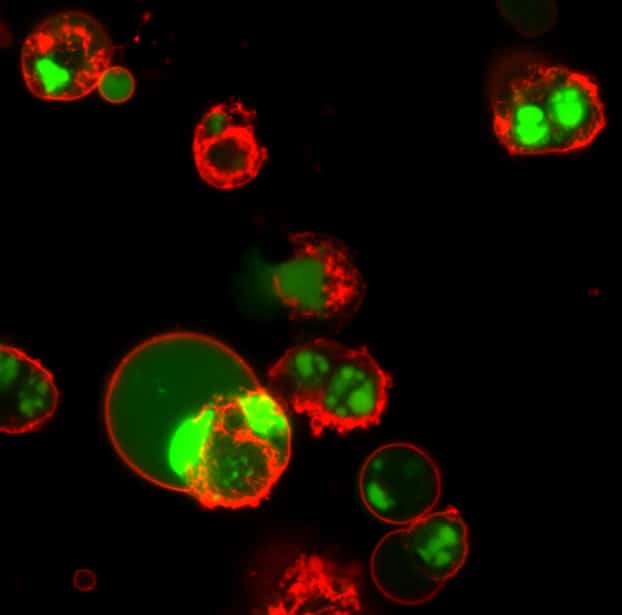
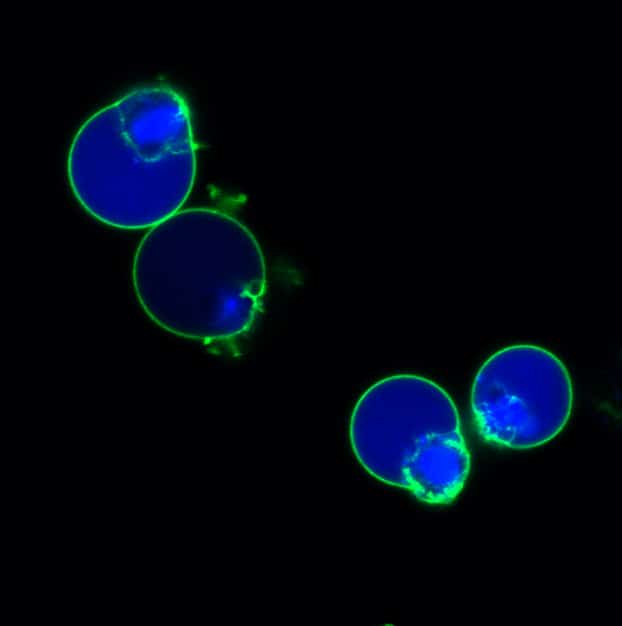
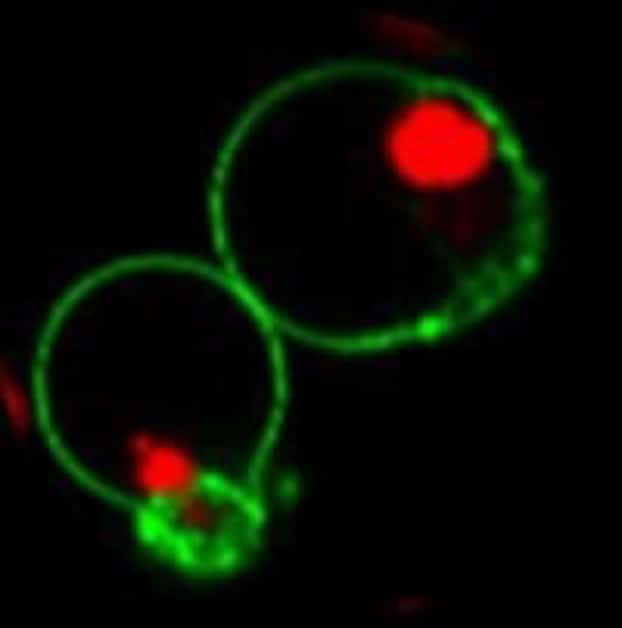
Available in Blue, Green, or Red Fluorescence
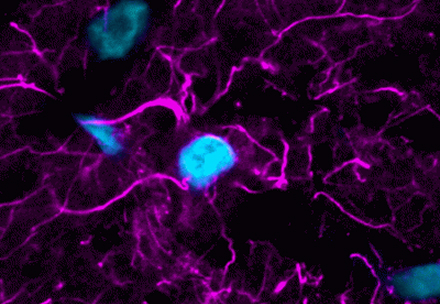
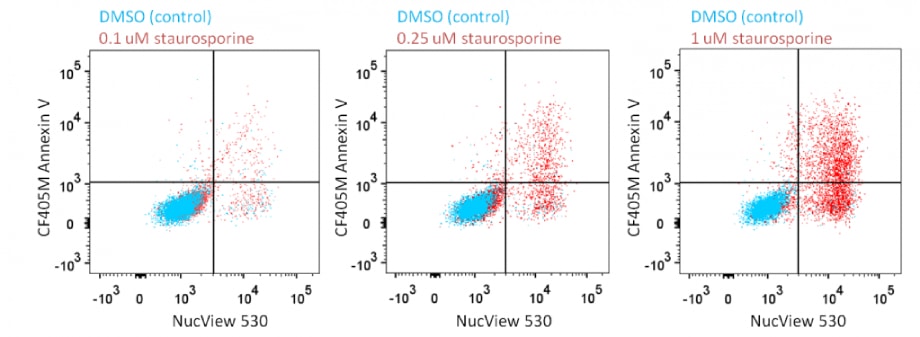
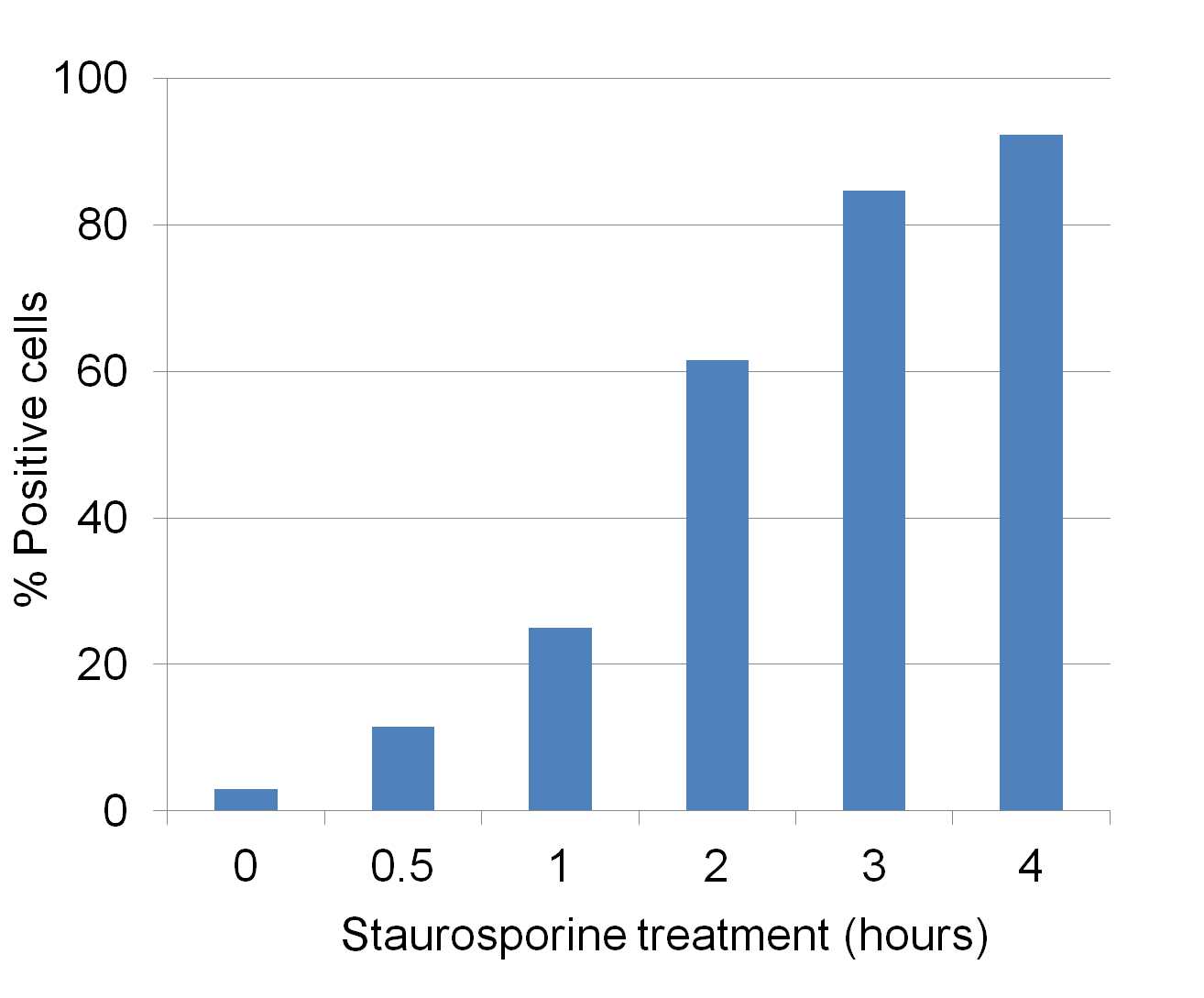
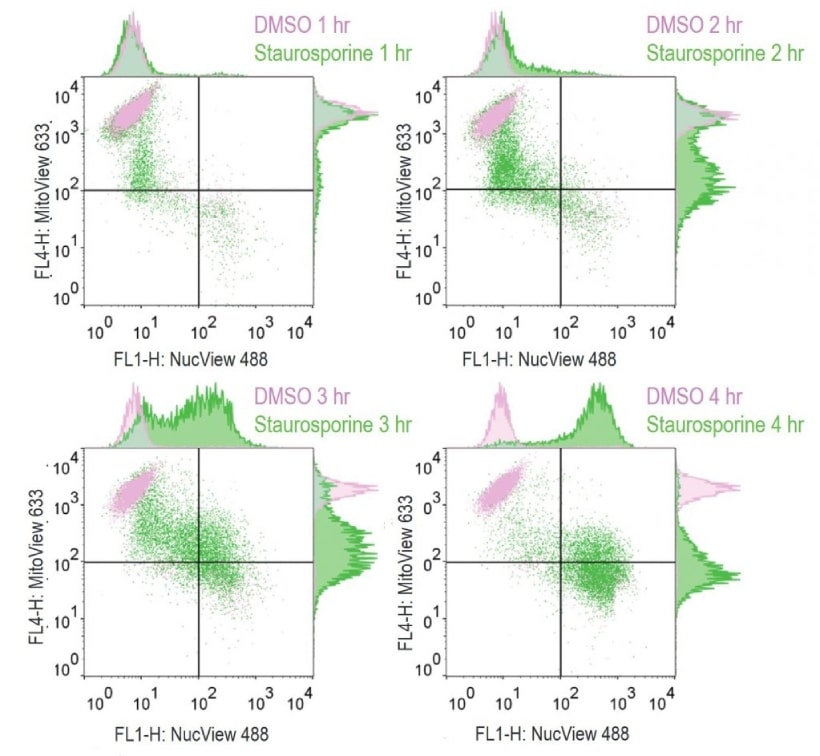
Analyze Apoptosis by Flow Cytometry in Live Cells
Dual Detection Kits
Explore assay kits that combine the advantage of NucView® real-time apoptosis analysis with other probes for monitoring necrosis, membrane potential, or other apoptosis markers.
- Dual Apoptosis Kit: NucView® 488 and Annexin V for co-detection of caspase cleavage and PS translocation.
- Dual Apoptosis Kit: NucView® 488 and MitoView™ 633 for co-detection of caspase cleavage and loss of mitochondrial potential.
- Apoptosis/Necrosis Kit: NucView® 488 and RedDot™2 for co-detection of caspase cleavage and loss of membrane integrity.
Full List of NucView® Substrates & Kits
NucView® Caspase-3 Substrates and Kits | Catalog No. | Features |
|---|---|---|
| NucView® 405 Caspase-3 Substrate, 1 mM in DMSO | 10405 | Blue fluorescence for flow cytometry in the Pacific Blue® channel or microscopy with the 405 nm laser |
| NucView® 405 Caspase-3 Substrate, 1 mM in PBS | 10407 | NucView® 405 substrate in PBS, for DMSO-sensitive cell types |
| NucView® 488 Caspase-3 Substrate, 1 mM in DMSO | 10402 | Green fluorescent substrate validated in more than 100 cell types and 200 publications |
| NucView® 488 Caspase-3 Substrate, 1 mM in PBS | 10403 | NucView® 488 substrate in PBS, for DMSO-sensitive cell types |
| NucView® 530 Caspase-3 Substrate, 1 mM in DMSO | 10406 | Orange fluorescence for microscopy in the Cy®3 channel or flow cytometry in the R-PE channel |
| NucView® 530 Caspase-3 Substrate, 1 mM in PBS | 10408 | NucView® 530 substrate in PBS, for DMSO-sensitive cell types |
| NucView® 488 and MitoView™ 633 Apotosis Detection Kit | 30062 | NucView® 488 and far-red fluorescent MitoView™ 633 for the Cy®5 channel |
| NucView® 488 and RedDot™ 2 Apoptosis & Necrosis Kit | 30072 | NucView® 488 and far-red dead cell DNA dye RedDot™ 2 for the Cy®5 channel |
| Dual Apoptosis Assay with NucView® 488 and CF®594 Annexin V | 30067 | NucView® 488 and red fluorescent Annexin V apoptosis probe |
| Dual Apoptosis Assay with NucView® 488 and CF®640R Annexin V | 30073 | NucView® 488 and far-red fluorescent Annexin V apoptosis probe |
The Gold Standard for Monitoring Apoptosis
Validated in Hundreds of Cell Types & Publications
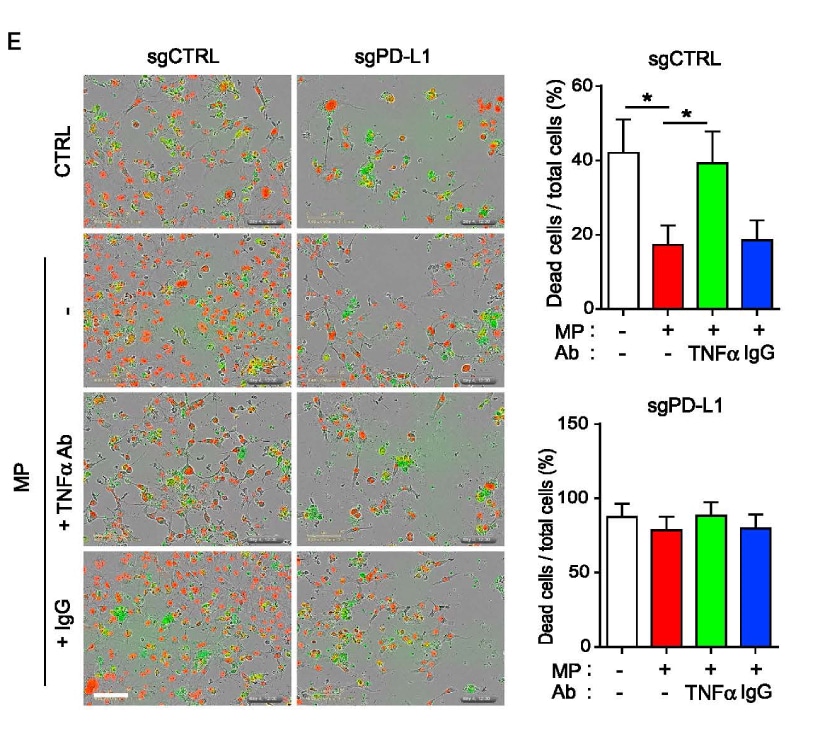
Highlighted Citation: NucView® 488 was used in a T cell-mediated tumor cell killing assay by Lim et al. in a study investigating the role of inflammation on PD-L1 stabilization. Results show T-cell mediated killing of PD-L1 expressing BT549 ductal carcinoma cells was no longer suppressed when treated with neutralizing TNF-α antibody. In addition, PD-L1 depleted cells did not respond to either cytokine enriched macrophage-conditioned medium (MP) or TNF-α antibody, suggesting that PD-L1 expression is required for TNF- α-mediated suppression of T cell activity.