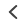Since the onset of the SARS-CoV-2 global pandemic in late 2019, mutated variants of concern (VOCs) have emerged that elicit the need for more sensitive surveillance and efficient detection. This is especially important for the Delta and Omicron variants, as these two are highly transmissible and contain multiple S gene mutations. Such factors underscore the importance for timely detection beyond the current genomic and Sanger sequencing methods that are typically both time consuming and costly.
In a May 2023 publication in Microbiology Spectrum, L Sun et al. developed a high-resolution melting (HRM®) assay to effectively detect SARS-CoV-2 variants without using time consuming and costly sequencing methods. This technique integrated multiple processes into a single-tube reaction and utilized Biotium’s EvaGreen® Dye to target eight key mutation sites, including those in the receptor binding domain (RBD) associated with enhanced infectivity and immune escape. Biotium’s EvaGreen® Dye was purposefully chosen by the researchers for its sensitivity, reliability, and cost-effectiveness. After analyzing 324 clinical samples of nasopharyngeal swabs and sputum through reverse transcription-quantitative PCR (qRT-PCR) and the multiplex HRM® method, both methods detected 48 samples positive for SARS-CoV-2, with strains of Delta, Omicron-BA.1, Omicron-BA.2, and Omicron-BA.5. The multiplex HRM® analysis had a limit of detection (LOD) lower than 10 copies/reaction for various targets and exhibited no cross-reactivity with other pathogens, achieving a robust 97.9% concordance rate with standard Sanger sequencing. The multiplex HRM® analysis clearly identified differentiated mutations in approximately two hours. The authors found that multiplex HRM® to be more reliable, economical, rapid, and expandable than sequencing for monitoring new viral variants as they emerge. With ever-evolving SARS-CoV-2 variants, this method may empower researchers and healthcare professionals with a convenient and faster alternative for timely diagnosis and surveillance.
Full Citation
Sun, L., Wang, L., Zhang, C. et al (2023). Rapid detection of predominant SARS-COV-2 variants using multiplex high-resolution melting analysis. Microbiology Spectrum (2023). https://journals.asm.org/doi/10.1128/spectrum.00055-23
HRM® is a registered trademark of Idaho Technologies, Inc./BioFire Defense, LLC