Content #1
Content #1
Content #1
Förster Resonance Energy Transfer (FRET) is an ideal tool to study conformational changes or binding interactions in biological systems due to its sensitivity to changes in distance between fluorophores on the nanometer scale. Two-color FRET, with one donor and one acceptor, is constrained to spatial resolution in one dimension. Thus the kinetic models that can be extrapolated from the data are limited.
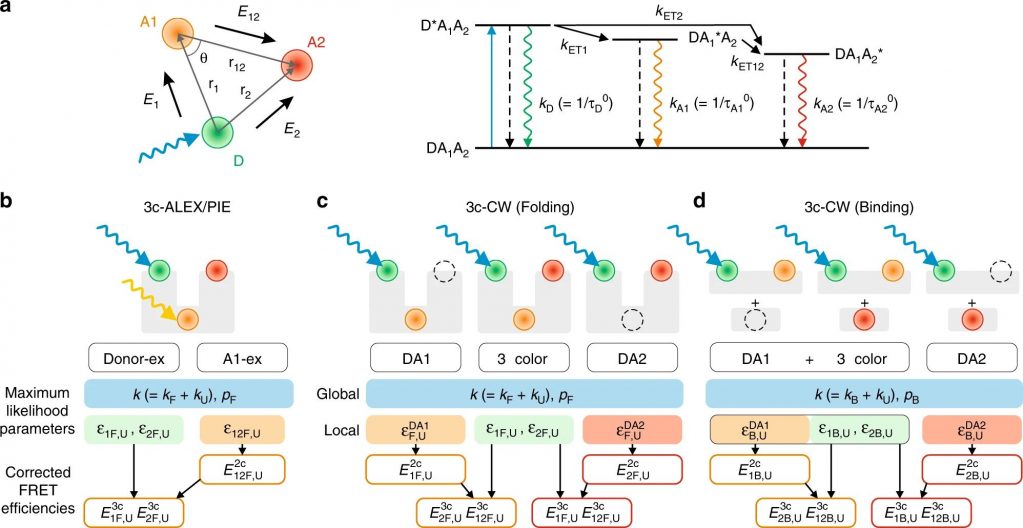
In a recent publication in Nature Communications, J. Yoo et al. developed a three-color FRET spectroscopic analysis using continuous-wave laser excitation to study fast molecular processes occurring on the microsecond to millisecond timescale. Three-color FRET uses a donor and two acceptor fluorophores to provide improved spatial resolution of molecular conformations of biological macromolecules. Three-color FRET is challenging because introducing additional acceptor excitation increases photophysical problems, including photoblinking, rapid photobleaching, and low quantum yield. The reactive fluorophores chosen to label the peptides, including CF®680R maleimide, are photostable and have a high quantum yield and thus overcome these challenges.
The authors tested the three-color FRET with two well characterized model systems: Protein folding of a de novo designed three helix bundle (alpha3D) and the binding of the N-terminal transactivation domain of the tumor suppressor protein p53 (TAD) with one of its binding partners (NCBD). The kinetics of folding and binding was monitored for both systems by following the donor and acceptors’ fluorescence intensity and were fit with simple two-state models. The authors conclude that folding and binding pathways can more accurately be characterized by three-color FRET than by a one-dimensional probe.
In another publication using three-color FRET Kim, JY and Chung, HS studied the transition states of the coupled folding and binding of TAD and NCBD. Transition states are stochastic and cannot be synchronized for ensemble measurement, but by using single molecule three-color FRET, the authors are able to obtain multidistance structural and kinetic information of individual transition states.
The authors demonstrate in these studies that single molecule three-color FRET can be useful in future studies of protein folding, binding kinetics as well as transition states of folding and disordered proteins.
Learn more about reactive CF® Dyes, GloMelt™ thermal protein stability assays, and Mix-n-Stain™ labeling kits.
Full citation
Yoo, J., Kim, JY., Louis, J.M. et al. Fast three-color single-molecule FRET using statistical inference. Nat Commun 11, 3336 (2020). https://doi.org/10.1038/s41467-020-17149-w
Kim, JY and Chung, HS, Disordered Proteins Follow Diverse Transition Paths as They Fold and Bind to a Partner. Science 368, 1253 (2020) https://doi.org/10.1126/science.aba3854