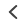RNA and RNA-based molecular machines participate in a range of important biological processes. In addition to functioning as messenger molecules between transcription and translation, they play roles in splicing, protein synthesis, and gene regulation. To perform these functions, RNA molecules need to be folded into compact, thermodynamically stable conformations. RNA chaperones, a class of intrinsically unstructured proteins, facilitate the folding process without a well-defined, three-dimensional structure. In a recent Nature Communications article, Holmstrom et al. address how RNA chaperones perform this function in a natively disordered state.
The authors use confocal single-molecule Förster resonance energy transfer (sm FRET) spectroscopy and molecular simulations to elucidate the mechanism of chaperone-assisted folding. Oligonucleotide hairpins labeled at the 5′ and 3′ ends with CF®660R (acceptor) and Cy®3b (donor) dyes were used to monitor stem-loop formation or ‘folding’ in the presence of the nucleocapsid domain (NCD) of the hepatitis C virus, an RNA chaperone. Their results show that NCD binds the hairpin with high affinity and acts as a flexible polycationic macromolecular counterion that accelerates nucleic acid folding by screening the repulsive negative charges along the phosphate backbone. While RNA chaperones employ a variety of mechanisms, the authors demonstrate that electrostatic charge screening serves as the primary factor underlying chaperone-assisted folding and propose its applicability to a wide range of nucleic acid targets in vivo.
Learn more about Biotium’s wide selection of CF® dyes and available reactive forms.
Full citation
Disordered RNA chaperones can enhance nucleic acid folding via local charge screening. Holmstrom ED, Liu Z, Nettels D, Best RB, Schuler B. Nat Commun. 2019;10(1):2453, doi: 10.1038/s41467-019-10356-0.