Normal light microscopy has inherent resolution limits preventing it from distinguishing certain intracellular structures. Methods like electron microscopy can be used for applications that require higher resolution but can be prohibitively expensive and require tedious sample preparation. In 2015, Chen, Tillberg, and Boyden of MIT described a method, now known as expansion microscopy (ExM), to enhance resolution by swelling samples instead of using higher resolution imaging instruments. Current ExM methods involve long, multi-step protocols. Cells are labeled with molecular “handles” such as antibody conjugates or reactive chemicals prone to quick hydrolysis in water, then perfused with a polymer gel that covalently links to the handles. Cellular material is then digested, and the gel expands, causing the attached labels to expand isotopically, allowing formerly nanoscale structures to be resolved using light microscopy. An ideal ExM protocol would be straightforward, provide ≥10-fold expansion, allow diverse labeling after expansion, and be compatible with a broad range of tissue types.
In their 2023 Nature Biotechnology article, Klimas et al. describe a novel ExM strategy named Magnify that meets the criteria for ideal ExM listed in the paragraph above. The key innovations of Magnify that lead to these improvements are a combined anchoring and gelling step, a novel gel formula, and homogenization with a hot denaturant-rich solution. Methacrolein, a classic small molecule fixative, was chosen to directly anchor biomolecules during gelation because it is stable in the gelling solution and fixes similarly to formaldehyde while also providing isopropenyl functional groups that assist during in situ gel polymerization. The authors also developed a novel hydrogel formula that is mechanically durable. They then used a panel of primary antibodies and Biotium CF® Dye-conjugated secondary antibodies, along with both pre- and post-expansion staining, to demonstrate the Magnify technique. To test for protein retention during homogenization and expansion, total sample protein was stained nonspecifically using a CF® Dye Succinimidyl Ester. Then, in order to delimit cell boundaries, glycocalyx was stained with CF® Dye conjugated WGA. The authors beautifully demonstrate Magnify using cryosections of mouse striatum, FFPE sections of several human tissues (Figure 1), stem cell-derived airway basal cells, and HEK-293FT cells. This relatively direct method provided ≥10-fold expansion in a wide range of tissues with various fixation methods, allowing imaging at ≈25 nm effective resolution on conventional optical microscopes or ≈15 nm using super-resolution optical fluctuation imaging. 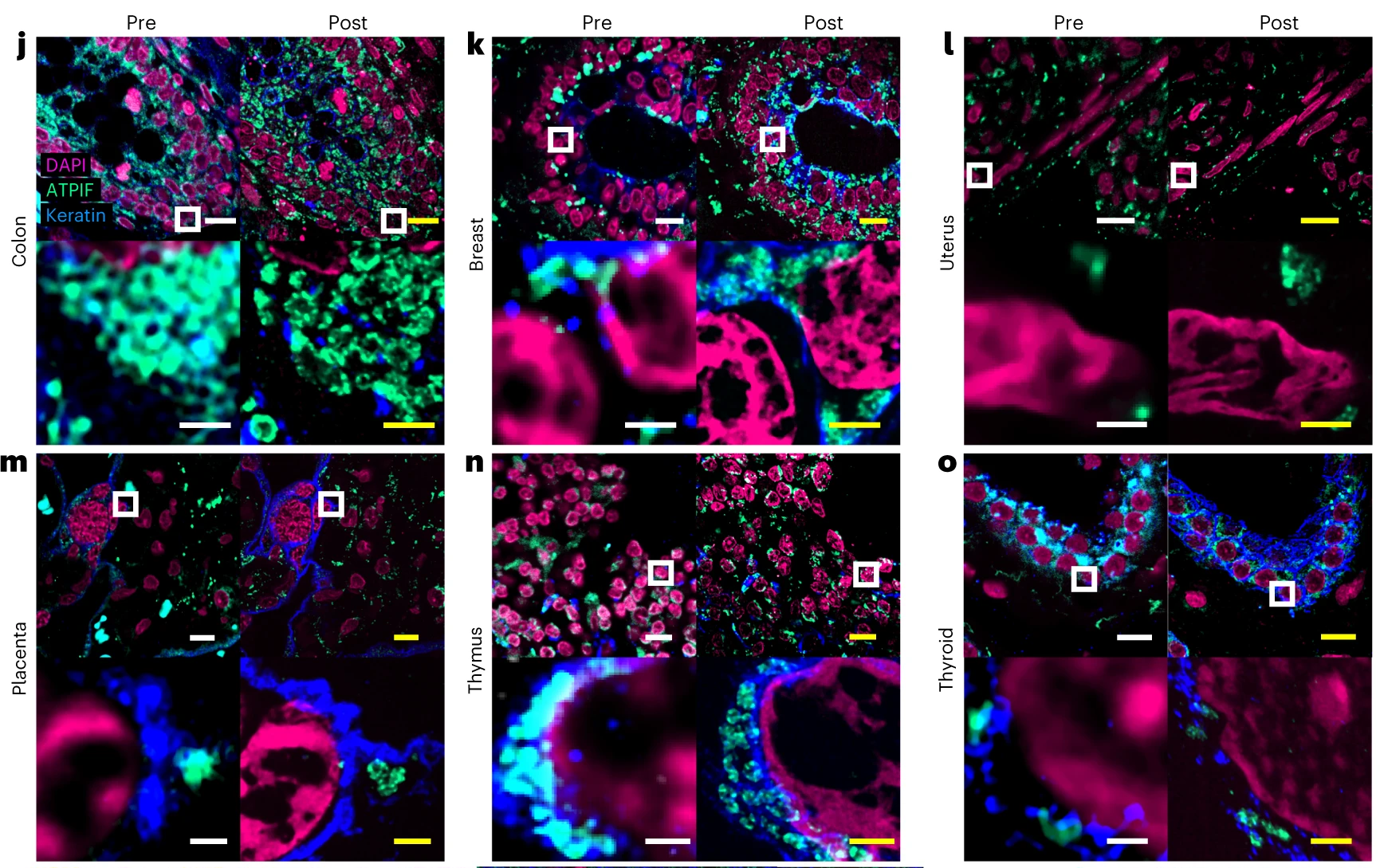 Figure 1. Validation of Magnify across diverse human FFPE samples. Tissues were imaged at ×40 (top left in each panel). Images were taken at ×60 and processed with SOFI (bottom left). The white box indicates the field of view (FOV) of the higher magnification images. The samples were then processed with the Magnify protocol, and the same FOVs were imaged post-expansion ×10 (top right) and ×40 (bottom right, projected over 4–17 z slices). Linear expansion factors (EF) post-expansion for each tissue were colon 8.85× (j), breast 9× (k), uterus 8× (l), placenta 8.75× (m), thymus 10.00× (n) and thyroid 10.59× (o). Scale bars in biological scale (yellow, post-expansion images). j–o, top, 10 μm; bottom, 1 μm; p–r, 5 μm. Credit: A. Klimas, et al. https://doi.org/10.1038/s41587-022-01546-1 reproduced under the Creative Commons license.
Figure 1. Validation of Magnify across diverse human FFPE samples. Tissues were imaged at ×40 (top left in each panel). Images were taken at ×60 and processed with SOFI (bottom left). The white box indicates the field of view (FOV) of the higher magnification images. The samples were then processed with the Magnify protocol, and the same FOVs were imaged post-expansion ×10 (top right) and ×40 (bottom right, projected over 4–17 z slices). Linear expansion factors (EF) post-expansion for each tissue were colon 8.85× (j), breast 9× (k), uterus 8× (l), placenta 8.75× (m), thymus 10.00× (n) and thyroid 10.59× (o). Scale bars in biological scale (yellow, post-expansion images). j–o, top, 10 μm; bottom, 1 μm; p–r, 5 μm. Credit: A. Klimas, et al. https://doi.org/10.1038/s41587-022-01546-1 reproduced under the Creative Commons license.
Featured videos from Klimas et, al. 2023
Learn more about Biotium’s high-performance CF® Dyes with multi-color flexibility, CF® Dye-conjugated secondary antibodies, and reagents for immunofluorescence microscopy.
Full Citation
Klimas, A., Gallagher, B. R., Wijesekara, P., et al. Magnify is a universal molecular anchoring strategy for expansion microscopy. Nat Biotechnol 1-12 (2023). https://doi.org/10.1038/s41587-022-01546-1