The Best Probes for Super-Resolution
Super-resolution microscopy encompasses a broad family of imaging techniques that push beyond the diffraction limit of traditional light microscopy, revealing the finer details of biological structures. These techniques rely on extremely precise control over the excitation, emission, and image acquisition of fluorescently labeled cells and tissues. Consequently, the quality and efficiency of super-resolution imaging relies heavily on the fluorescent properties of the probe. Biotium’s CF® Dyes have met this demand with industry-leading brightness and photochemical switching properties over other commercially available fluorescent probes. Moreover, CF® Dyes have been published in dozens of studies for a wide range of super-resolution applications.
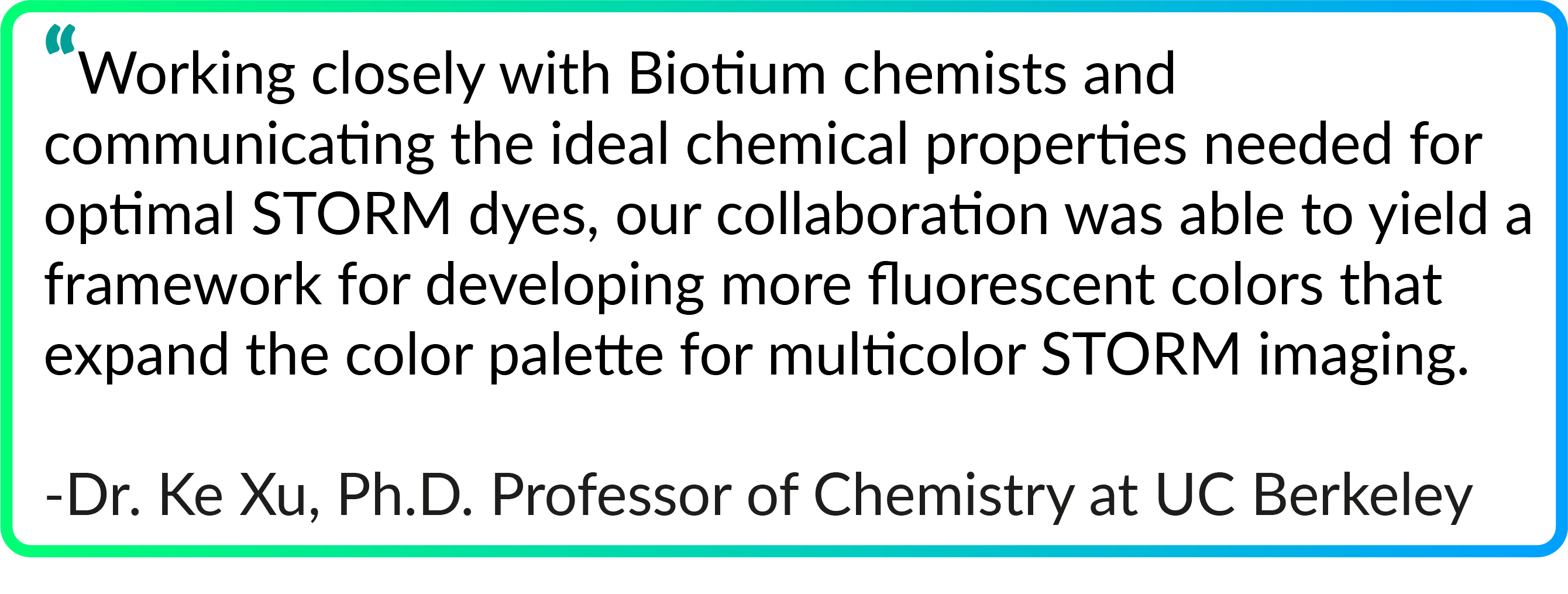
Superior Performance for STORM Imaging
Several CF® Dyes are specifically developed and validated for optimal performance in Stochastic Optical Reconstruction Microscopy (STORM). This includes Biotium’s red-excited CF® Dyes such as CF®647, CF®660C, and CF®680 which display superior brightness and unique photoswitching properties that are ideal for STORM. However, while STORM imaging commonly relies on photoswitchable red-excited dyes, finding comparable dyes for other excitation sources to produce quality multi-color STORM images remains a challenge. In response, Biotium has developed several green-excited dyes developed for optimal performance in STORM. Specifically, green-excited CF®583R and CF®597R were developed in collaboration with UC Berkeley and STORM scientist Ke Xu, PhD for exceptional performance in multi-color STORM imaging (Figures 1 and 2). In addition, green-excited CF®568 has also demonstrated better performance than Cy3b (Figure 3). CF®594ST is Biotium’s unique version of CF®594 developed specifically for STORM. See below for a full list of validated CF® Dyes for STORM and available products.
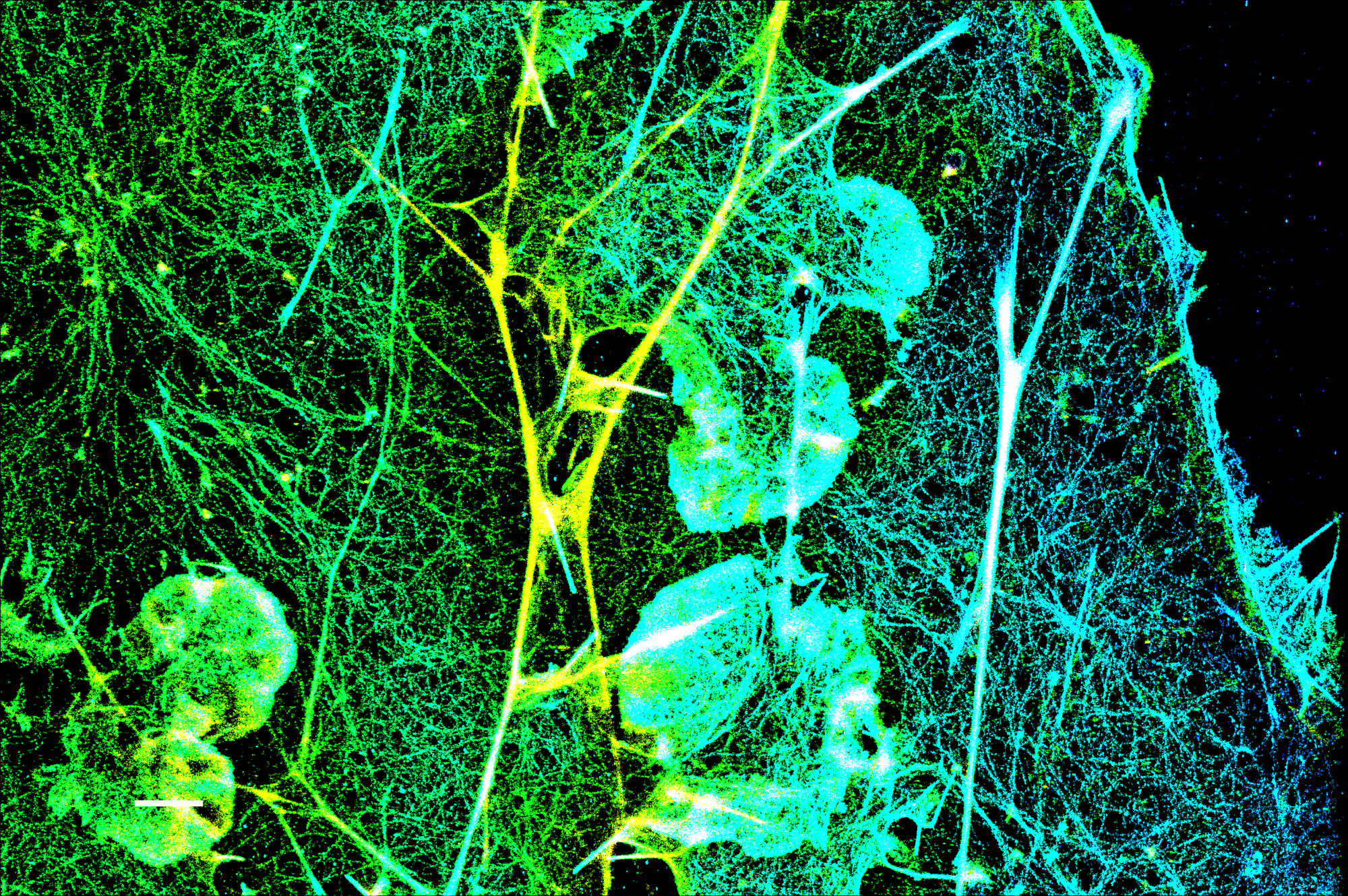
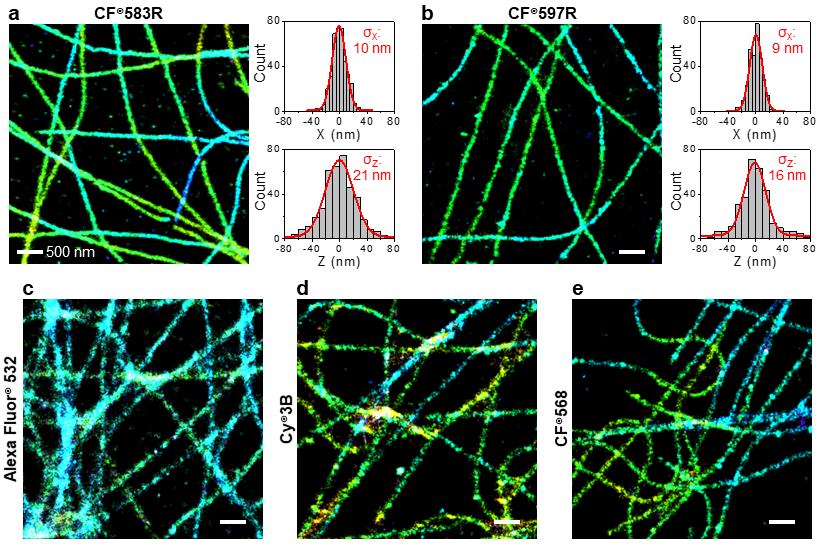
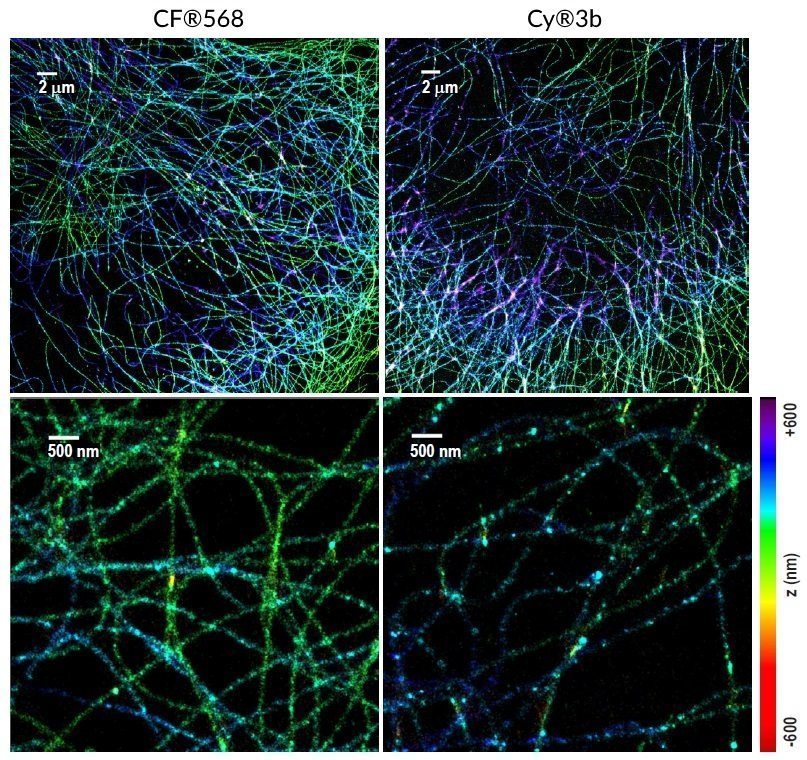
CF® Dyes Validated for STED and FLIM Imaging by STELLARIS 8 STED Confocal Microscope
CF® Dyes were validated for high-quality super-resolution imaging of microtubules by Leica’s STELLARIS 8 STED FALCON platform. See the table on the right for fluorescent lifetime data for several CF® Dyes.
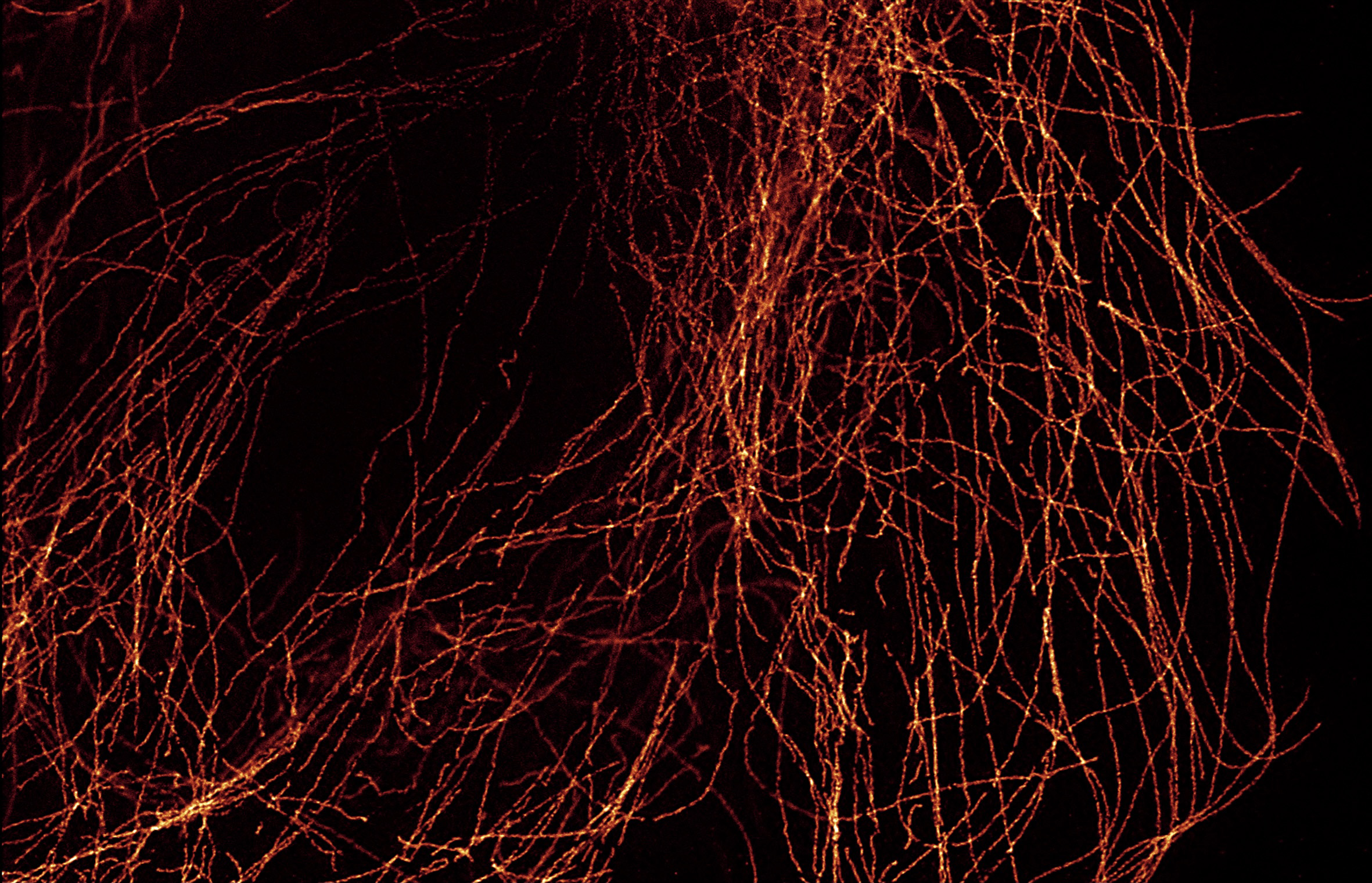
CF® Dye Fluorescence Lifetime Data
| Dye | τ (ns) /free acid in PBS pH 7.4, ε (ns) | τ (ns) /S.Ab§ |
|---|---|---|
| CF®405S | 3.88 ± 0.05 | – |
| CF®488A | 4.11 ± 0.05 | 1.705 |
| CF®568 | 3.66 ± 0.05 | 1.539 |
| CF®594 | – | 1.746 |
| CF®633 | 3.39* (in water) | 3* |
| CF®640R | 2.38 ± 0.05 | 1.557 |
| CF®647 | 1.07 ± 0.05 | 1.195 |
| CF®680 | 1.23 ± 0.05 | 1.277 |
| CF®680R | 1.22 ± 0.05 | 1.6 |
| CF®750 | 0.58 ± 0.05 | 0.636 |
| CF®790 | 0.39 ± 0.05 | 0.54 |
Measurements were made on a Stellaris 8 STED FALCON microscope courtesy Leica Microsystems, Germany. § Fluorescence lifetime measurements of CF® Dye labeled anti-mouse secondary antibodies used for immunostaining microtubules in U2OS using mouse anti-tubulin (DM1a) and mounted in ProLong™ Diamond. *Lifetime data obtained via customer communication under different experimental conditions and imaging setup.
CF® Dye Products Useful for Super-Resolution Microscopy
Antibody Labeling Kits
Our Mix-n-Stain™ STORM CF® Dye Antibody Labeling Kits allow you to label 50 ug of your antibody with one of Biotium’s STORM CF® Dyes to produce an antibody conjugate with a low 1-2.5 DOL (degree of labeling) optimal for super-resolution STORM (Stochastic Optical Reconstruction Microscopy). Labeling takes just 30 minutes, with minimal hands-on time and no purification. Click here to learn more about Biotium’s antibody labeling kits.
- Choice of 8 CF® Dye colors ideal for STORM
- CF®583R and CF®597R are novel green-excited STORM dyes
- Labeling optimized to provide low 1-2.5 DOL
- Less than 30 seconds of hands-on time
- 30 minutes total reaction time
- No purification, 100% recovery
Reactive CF® Dyes
Developing a custom probe for super-resolution applications? Biotium offers CF® Dyes in a wide array of reactive chemistries to suit your research needs.
Single-Label Secondary Antibody Conjugates for STORM
Secondary antibodies with a low degree of labeling (DOL, or number of dye molecules per antibody molecule) have been reported to be optimal for STORM (Bittel et al. 2015). Biotium offers single-label secondary antibody conjugates with an average DOL of one for STORM applications.
Probes for Cytoskeleton and Other Cellular Structures
Biotium offers a wide variety of CF® Dye bioconjugates used for labeling specific cellular structures.
- Phalloidin conjugates for F-actin labeling available in 16 CF® Dyes. CF®647 and CF®680 phalloidins ideal for STORM imaging.
- Con A, WGA, and PNA lectin CF® Dye conjugates for labeling glycoproteins and cellular surfaces
- Streptavidin & biotinylated CF® Dye conjugates
- CF® Dye nucleotides and probes for apoptosis, endocytosis, and other cellular processes also available.
Learn more about CF® Dyes for Super-Resolution Imaging:
CF® Dyes Validated for Super-Resolution Microscopy
| Dye | Abs/Em (nm) | STORM | STED | SIM | 2-Photon | TIRF | Other Applications | Features |
|---|---|---|---|---|---|---|---|---|
| CF®405S | 411/431 | ✓ | ✓ | • Brighter than Alexa Fluor® 405 | ||||
| CF®405M | 416/452 | ✓ | ✓ | ✓ | • More photostable than Pacific Blue™ • Excellent choice for SIM imaging |
|||
| CF®440 | 433/512 | ✓ | • More photostable than spectrally similar dyes | |||||
| CF®488A | 490/516 | ✓ | ✓ | ✓ | ✓ | ✓ | DNA-PAINT | • Less non-specific binding than Alexa Fluor® 488 |
| CF®498 | 498/519 | ✓ | • Less non-specific binding than Alexa Fluor® 488 • Use with CF®568/CF®583R and CF®647/Alexa Fluor®647 for multicolor STORM |
|||||
| CF®505 | 505/519 | ✓ | • Identical to ATTO 488 | |||||
| CF®535ST | 535/569 | ✓ | • Orange dye designed specifically for STORM imaging | |||||
| CF®555 | 554/568 | ✓1 | ✓1 | • Brighter and more photostable than Cy®3 • Less non-specific binding than Alexa Fluor® 555 |
||||
| CF®568 | 562/584 | ✓1 | ✓ | ✓ | ✓ | • Yields much brighter conjugates than Alexa Fluor® 568 • Outperforms Cy®3b in STORM • Pairs well with CF®647 and CF®680 for multi-color STORM – see publication |
||
| CF®583R | 585/609 | ✓1 | • One of two top-performing dyes specifically designed for STORM with green laser (also see CF®597R) – see publication |
|||||
| CF®594 | 593/615 | ✓ | ✓ | • Significantly brighter than Alexa Fluor® 594 and Texas Red® • Extremely photostable |
||||
| CF®597R | 597/619 | ✓1 | • Deep-red fluorescent dye designed specifically for STORM • Top-performing dye specifically designed for STORM with green laser (also see CF®583R) – see publication |
|||||
| CF®633 | 629/650 | ✓ | FIONA, gSHRImp, SMT | • Significantly brighter than spectrally similar far-red dyes • Far more photostable than Alexa Fluor® 647 |
||||
| CF®640R | 642/663 | ✓ | ✓ | ✓ | ✓ | FLImP | • Offers improved brightness and photostability over ATTO 647N and spectrally similar dyes |
|
| CF®647 | 652/668 | ✓1 | • Spectrally similar to Cy®5 and Alexa Fluor® 647 • Pairs well with CF®568 for multi-color STORM • The best far-red dye for demixing-based multi-color (d)STORM imaging when paired with CF®680 – see publication |
|||||
| CF®660R | 662/682 | SMLM, DNA-PAINT | • Much brighter than Alexa Fluor® 660 • The most photostable 660 nm dye • Validated for use with DNA-PAINT SMLM – see publication |
|||||
| CF®660C | 667/685 | ✓1 | MINFLUX | • Much brighter and more photostable than Alexa Fluor® 660 • Ideal for long high-intensity 3D (d)STORM image acquisitions with minimal photobleaching – see publication |
||||
| CF®680 | 681/698 | ✓1 | Dual-color 3D SMLM, MINFLUX | • The brightest among spectrally similar 680 nm dyes • Pairs well with CF®568 for multi-color STORM • The best near-IR dye for demixing-based multi-color (d)STORM imaging when paired with CF®647 – see publication |
||||
| CF®680R | 680/701 | ✓1 | ✓ | ✓ | Single-molecule spectroscopy, SMT | • The most photostable 680 nm dye • Suitable for labeling nucleic acids and small biomolecules |
||
| CF®750 | 755/779 | ✓ | • Exceptionally bright and photostable near-IR dye • Patented pegylated dye for superior performance |
1 Dye was validated in multi-color STORM experiments
FLImP: Fluorophore localization imaging with photobleaching; SIM: Structured illumination microscopy; STED: Stimulated emission depletion; STORM: Stochastical optical reconstruction microscopy; TIRF: Total internal reflection fluorescence; FIONA: Fluorescence imaging with one-nanometer accuracy; ExM: Expansion microscopy; SMT: Single-molecule tracking; SMLM: Single-molecule localization microscopy.
References for CF® Dyes in Super-Resolution Microscopy and Other Specialized Applications
| Dye | Abs/Em (nm) | Applications | References |
|---|---|---|---|
| CF®405S | 404/431 | SIM | Demmerle, J. et al. (2017). Strategic and practical guidelines for successful structured illumination microscopy. Nature Protocols 12, 988–1010. (SIM) Essig, K. et al. (2017). Roquin Suppresses the PI3K-mTOR Signaling Pathway to Inhibit T Helper Cell Differentiation and Conversion of Treg to Tfr Cells. Volume 47, Issue 6, p1067–1082.e12 (SIM) |
| CF®405M | 408/452 | SIM STED 2-photon | Demmerle, J. et al. (2017). Strategic and practical guidelines for successful structured illumination microscopy. Nature Protocols 12, 988–1010. (SIM) Kraus, F. et al. (2017) Quantitative 3D structured illumination microscopy of nuclear structures.Nat Protoc 12, 1011-1028, doi:nprot.2017.020 [pii] (SIM) Markaki, Y. et al. (2013). Fluorescence In Situ Hybridization Applications for Super-Resolution 3D Structured Illumination Microscopy. Methods Mol Biol 950, 43-64. (SIM) Miron, E. et. al. (2016). In Vivo and In Situ Replication Labeling Methods for Super-resolution Structured Illumination Microscopy of Chromosome Territories and Chromatin Domains. in Mark C. Leake (ed.), Chromosome Architecture: Methods and Protocols, Methods in Molecular Biology, vol. 1431, 127-140. (SIM) Ohgomori, T. et al. (2017). Differential activation of neuronal and glial STAT3 in the spinal cord of the SOD1G93A mouse model of amyotrophic lateral sclerosis.Eur J Neurosci 46, 2001-2014, doi:10.1111/ejn.13650 (SIM) Zhang, R. et al. (2019). The mechanisms of dynamin-actin interaction. bioRxiv doi: https://doi.org/10.1101/586461 (STED) Kim, YR et al. (2020). Neutrophils Return to Bloodstream Through the Brain Blood Vessel After Crosstalk With Microglia During LPS-Induced Neuroinflammation. Front. Cell Dev. Biol. (2-photon) |
| CF®488A | 490/515 | STED STORM TIRF 2-photon | Angelov, B. & Angelova, A. (2017). Nanoscale clustering of the neurotrophin receptor TrkB revealed by super-resolution STED microscopy. Nanoscale 9, 9797-9804, doi:10.1039/c7nr03454g. (STED) Mercier, L. et al. (2016). In vivo imaging of skeletal muscle in mice highlights muscle defects in a model of myotubular myopathy. Intravital. 5(1), e1168553. (2-photon) Zanetti-Domingues, L.C. et al. (2013). Hydrophobic Fluorescent Probes Introduce Artifacts into Single Molecule Tracking Experiments Due to Non-Specific Binding. PLoS ONE 8(9): e74200. (TIRF) Collaborator communication (CF®488A for STORM using OxEA buffer); for more information contact Tech Support. |
| CF®535ST | 535/568 | STORM | Collaborator communication; for more information contact Tech Support. |
| CF®555 | 555/565 | Multicolor STORM | Lehmann, M. et al. (2015). Novel organic dyes for multicolor localization-based super-resolution microscopy. J Biophotonics DOI 10.1002/jbio.201500119 |
| CF®568 | 562/583 | Multicolor STORM SIM TIRF STED | Gong, Y.-N. et al. (2017). Biological events and molecular signaling following MLKL activation during necroptosis. Cell Cycle, 1-13. doi:10.1080/15384101.2017.1371889 (STORM) Gorur, A. et al. (2017). COPII-coated membranes function as transport carriers of intracellular procollagen I. J Cell Biol 216, 1745-1759. doi:10.1083/jcb.201702135 (STORM) Haas, K. et al. (2018). Single-molecule localization microscopy reveals molecular transactions during RAD51 filament assembly at cellular DNA damage sites. Nucleic Acids Research, Volume 46(5), 2398–2416. (STORM) Heller, J. (2017). Exploring Nanoscale Organisation of Synapses with Super-Resolution Microscopy. OM&P 3, 48-58, doi:doi:10.20388/omp2017.002.0045 (STORM) Jorgans, D.M. et al. (2017). Deep nuclear invaginations are linked to cytoskeletal filaments – integrated bioimaging of epithelial cells in 3D culture. J Cell Sci 2017 130: 177-189. doi: 10.1242/jcs.190967 (STORM) Karanasios, E. et al. (2016). Autophagy initiation by ULK complex assembly on ER tubulovesicular regions marked by ATG9 vesicles.Nature Communications 7: 12420. DOI: 10.1038/ncomms12420 (STORM) Kraus, F. et al. (2017) Quantitative 3D structured illumination microscopy of nuclear structures.Nat Protoc 12, 1011-1028, doi:nprot.2017.020 [pii] (SIM) Lehmann, M. et al. (2015). Novel organic dyes for multicolor localization-based super-resolution microscopy.J Biophotonics DOI 10.1002/jbio.201500119 (STORM) Lim, A. et al. (2017). Two kinesins drive anterograde neuropeptide transport.Mol Biol Cell, doi:mbc.E16-12-0820 [pii] (SIM) Turkowyd, B. et al. (2016). From single molecules to life: microscopy at the nanoscale. Anal Bioanal Chem DOI 10.1007/s00216-016-9781-8 (STORM) Zanetti-Domingues, L.C. et al. (2013). Hydrophobic Fluorescent Probes Introduce Artifacts into Single Molecule Tracking Experiments Due to Non-Specific Binding. PLoS ONE 8(9): e74200. (TIRF) Zhang, M. et al. (2015).Translocation of interleukin-1β into a vesicle intermediate in autophagy-mediated secretion. eLife 2015;10.7554/eLife.11205 (STORM) Catsburg, L. A. et al. (2021) Dynamics and nanoscale organization of the postsynaptic endocytic zone at 1 excitatory synapses. BioRxiv 2021.02.18.431766 (STED) |
| CF®583R | 585/609 | Multicolor STORM | Wang, B. et al. (2021). Transforming Rhodamine Dyes for (d)STORM Super-Resolution Microscopy via 1,3-Disubstituted Imidazolium Substitution. Angewandte Chemie International Edition, e202113612. |
| CF®594 | 593/614 | 2-photon STED | Wagner, M.C. et al. (2016). Mechanism of increased clearance of glycated albumin by proximal tubule cells. Am J Physiol Renal Physiol 310, F1089–F1102. (2-photon) Collaboration with Leica using the STELLARIS STED microscope (STED); for more information contact Tech Support. |
| CF®594ST | 593/614 | STORM | Collaborator communication; for more information contact Tech Support. Please note: CF®594ST is a unique dye designed specifically for STORM. Our original CF®594 dye is not suitable for STORM. |
| CF®597R | 597/619 | Multicolor STORM | Wang, B. et al. (2021). Transforming Rhodamine Dyes for (d)STORM Super-Resolution Microscopy via 1,3-Disubstituted Imidazolium Substitution. Angewandte Chemie International Edition, e202113612. |
| CF®633 | 630/650 | FIONA gSHRImP Single-molecule tracking (SMT) TIRF | Bosch, P.J. et al. (2014).Evaluation of fluorophores to label SNAP-tag fused proteins for multicolor single-molecule tracking microscopy in live cells. Biophys J 107, 803-814. (TIRF) Huang, T. et al. (2018). Simultaneous Multicolor Single-Molecule Tracking with Single-Laser Excitation via Spectral Imaging. Biophysical Journal 114, 301–310. (SMT) Kim, H.J., and Selvin, P.R. (2013). Fluorescence Imaging with One Nanometer Accuracy.SpringerReference Encyclopedia of Biophysics. (FIONA) Simonson, P. D. et al. (2011). Single-molecule-based super-resolution images in the presence of multiple fluorophores.Nano Lett 11, 5090-5096. DOI:10.1021/nl203560r (gSHRImP) Zanetti-Domingues, L.C. et al. (2013). Hydrophobic Fluorescent Probes Introduce Artifacts into Single Molecule Tracking Experiments Due to Non-Specific Binding. PLoS ONE 8(9): e74200. (TIRF) |
| CF®640R | 642/662 | FLImP SIM TIRF STED | Bosch, P. J. et al. (2014).Evaluation of fluorophores to label SNAP-tag fused proteins for multicolor single-molecule tracking microscopy in live cells. Biophys J 107, 803-814. (TIRF) Loh, L. N. (2017). Dissecting Bacterial Cell Wall Entry and Signaling in Eukaryotic Cells: an Actin-Dependent Pathway Parallels Platelet-Activating Factor Receptor-Mediated Endocytosis.MBio 8, doi:mBio.02030-16 [pii] (SIM) Martin-Fernandez, M. L. et al. (2013). A ‘pocket guide’ to total internal reflection fluorescence.J Microsc 252, 16-22. (TIRF) Needham, S.R. et al. (2015). Determining the geometry of oligomers of the human epidermal growth factor family on cells with <10 nm resolution. Biochem Soc Trans 43, 309-314. (FLImP) Needham, S.R. et al. (2016). EGFR oligomerization organizes kinase-active dimers into competent signalling platforms. Nat Commun 7, 13307. doi:ncomms13307 (FLImP) Zanetti-Domingues, L.C. et al. (2013). Hydrophobic Fluorescent Probes Introduce Artifacts into Single Molecule Tracking Experiments Due to Non-Specific Binding.PLoS ONE 8(9): e74200. (TIRF) Zanetti-Domingues, L.C. et al. (2015). Determining the geometry of oligomers of the human epidermal growth factor family on cells with 7 nm resolution. Prog Biophys Mol Biol 118, 139-152, doi:S0079-6107(15)00047-4 (FLImP) Zhang, R. et al. (2017). Structural insight into TPX2-stimulated microtubule assembly. eLife 2017;6:e30959. (TIRF) Collaboration with Leica using the STELLARIS STED microscope (STED); for more information contact Tech Support. |
| CF®647 | 650/665 | Multicolor STORM | Gong, Y.-N. et al. (2017). Biological events and molecular signaling following MLKL activation during necroptosis. Cell Cycle, 1-13. doi:10.1080/15384101.2017.1371889 Lehmann, M. et al. (2015). Novel organic dyes for multicolor localization-based super-resolution microscopy.J Biophotonics DOI 10.1002/jbio.201500119 Olivier, N. et al. (2013). Simple buffers for 3D STORM microscopy.Biomed Opt Express 4, 885-899. Turkowyd, B. et al. (2016). From single molecules to life: microscopy at the nanoscale.Anal Bioanal Chem DOI 10.1007/s00216-016-9781-8 |
| CF®660C | 667/685 | Multicolor STORM | Gu, L., Li, Y., Zhang, S. et al. Molecular-scale axial localization by repetitive optical selective exposure. Nat Methods 18, 369–373 (2021). https://doi.org/10.1038/s41592-021-01099-2 Turkowyd, B. et al. (2016). From single molecules to life: microscopy at the nanoscale. Anal Bioanal Chem DOI 10.1007/s00216-016-9781-8 Zhang, Z. et al. (2015). Ultrahigh-throughput single-molecule spectroscopy and spectrally resolved super-resolution microscopy. Nature Methods doi:10.1038/nmeth.3528 |
| CF®680 | 681/698 | Dual-color 3D SMLM Multicolor STORM | Früh, S.M. et al. (2015). Molecular architecture of native fibronectin fibrils. Nature Communications 6, 7275. (STORM) Glebov, O. O. et al. (2017).Nanoscale Structural Plasticity of the Active Zone Matrix Modulates Presynaptic Function. Cell Rep 18, 2715-2728. doi:S2211-1247(17)30279-6 [pii] (STORM) Gorur, A. et al. (2017). COPII-coated membranes function as transport carriers of intracellular procollagen I. J Cell Biol 216, 1745-1759. doi:10.1083/jcb.201702135 (STORM) Lehmann, M. et al. (2015). Novel organic dyes for multicolor localization-based super-resolution microscopy.J Biophotonics DOI 10.1002/jbio.201500119 (STORM) Platonova, E. et al. (2015). A Simple Method for GFP- and RFP-based Dual Color Single-Molecule Localization Microscopy. ACS Chem. Biol.10(6),1411-1416. (STORM) Platonova, E. et al. (2015). Single-molecule microscopy of molecules tagged with GFP or mRFP derivatives in mammalian cells using nanobody binders.Methods doi: http://dx.doi.org/10.1016/j.ymeth.2015.06.018 (STORM) Salvador-Gallego, R. et al. (2016). Bax assembly into rings and arcs in apoptotic mitochondria is linked to membrane pores. EMBO J 35, 389-401. doi:embj.201593384 (STORM) Shrestha, R. L. et al. (2017). Aurora-B kinase pathway controls the lateral to end-on conversion of kinetochore-microtubule attachments in human cells. Nat Commun 8, 150. doi:10.1038/s41467-017-00209-z (STORM) Thiele, J. C. et al. (2022). Isotropic three-dimensional dual-color super-resolution microscopy with metal-induced energy transfer. Sci. Adv. 8, eabo2506, doi:10.1126/sciadv.abo2506 (MIET-SMLM) Turkowyd, B. et al. (2016). From single molecules to life: microscopy at the nanoscale.Anal Bioanal Chem DOI 10.1007/s00216-016-9781-8 (SMLM) Winterflood, C.M. et al. (2015). Dual-Color 3D Superresolution Microscopy by Combined Spectral-Demixing and Biplane Imaging. Biophys J. 109, 3-6. (SMLM) Zhang, Z. et al. (2015). Ultrahigh-throughput single-molecule spectroscopy and spectrally resolved super-resolution microscopy.Nature Methods doi:10.1038/nmeth.3528 (STORM) |
| CF®680R | 680/701 | Multicolor STORM Single-molecule spectroscopy Single-molecule tracking (SMT) STED 2-photon | Conley, G. et al. (2017). Jamming and overpacking fuzzy microgels: Deformation, interpenetration, and compression. Science Advances 3(10), e1700969 DOI:10.1126/sciadv.1700969 (STORM) Görlitz, F. et al. (2014). A STED Microscope Designed for Routine Biomedical Applications. Progress Electromagnetics Res 147, 57-68. (STED) Huang, T. et al. (2018). Simultaneous Multicolor Single-Molecule Tracking with Single-Laser Excitation via Spectral Imaging. Biophysical Journal 114, 301–310. (SMT) König, I. et al. (2015). Single-molecule spectroscopy of protein conformational dynamics in live eukaryotic cells. Nature Methods doi:10.1038/nmeth.3475 (Single molecule spec) Moran, I. et al. (2018). Memory B cells are reactivated in subcapsular proliferative foci of lymph nodes. Nature Communications 9:3372. (2-photon) Winter, F. R. et al. (2017). Multicolour nanoscopy of fixed and living cells with a single STED beam and hyperspectral detection.Sci Rep 7, 46492. doi:srep46492 [pii]10.1038/srep46492 (STED) |
| CF®750 | 755/777 | STORM | Collaborator communication; for more information contact Tech Support. |


