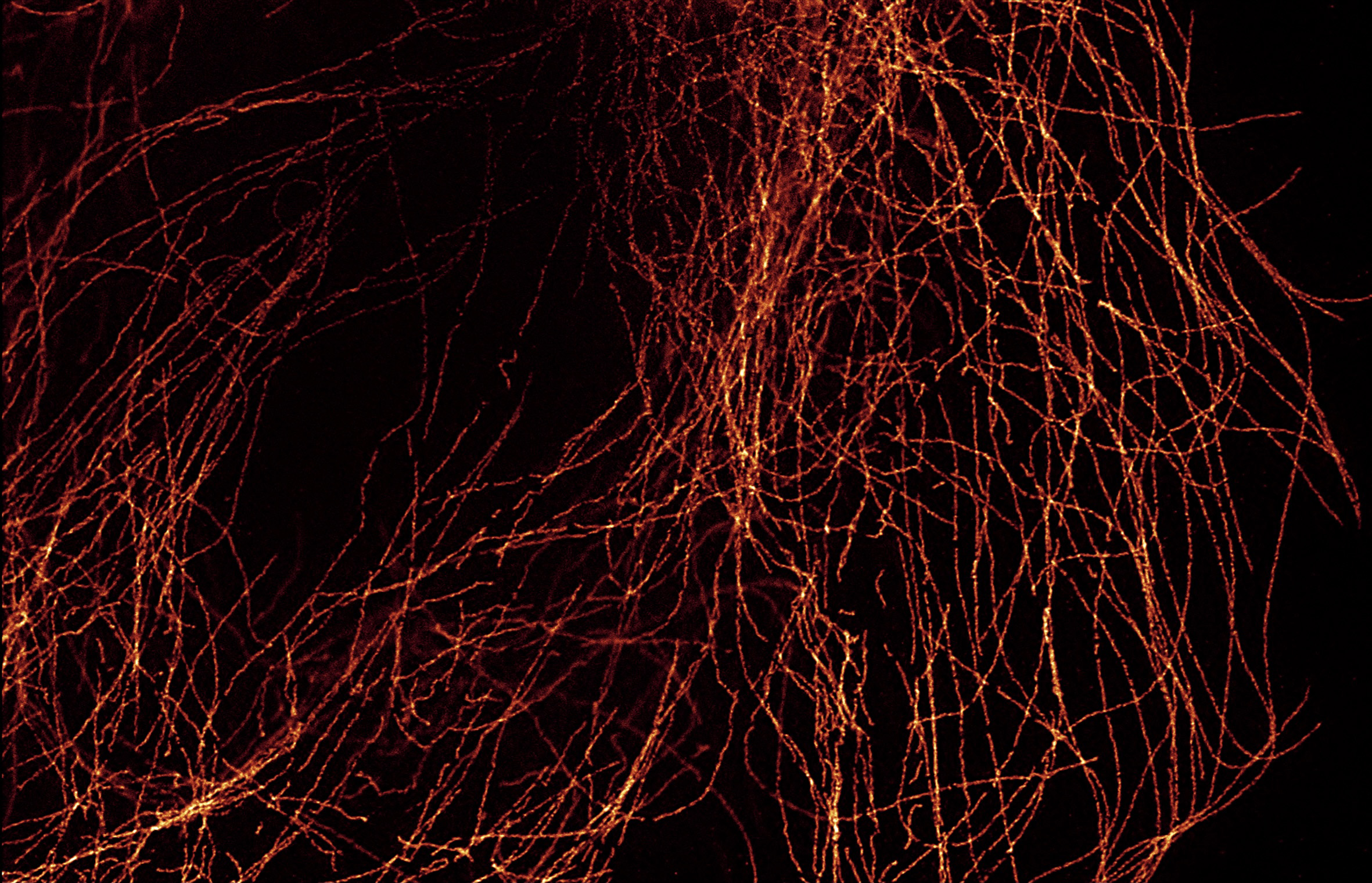
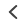Super resolution microscopy has revolutionized our understanding of biology by shedding light on the molecular world beyond the diffraction limit. Traditional light microscopy is limited to about 200 nm resolution under ideal circumstances, where resolution is roughly one-half the wavelength of emitted light. Everything smaller than this limit appears as a diffraction limited spot.
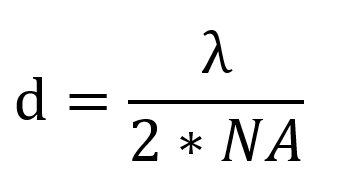
Super resolution microscopy has revealed previously unknown structures in biology and shown us the dynamic behavior of individual molecules. Super resolution microscopy uses physical tricks to see beyond the diffraction limit, including illuminating samples with a shifting pattern, depleting fluorescence at the edge of the excitation beam, and imaging single molecules. These techniques are constantly improving with advances in dye creation, acquisition speed, and reducing phototoxicity.
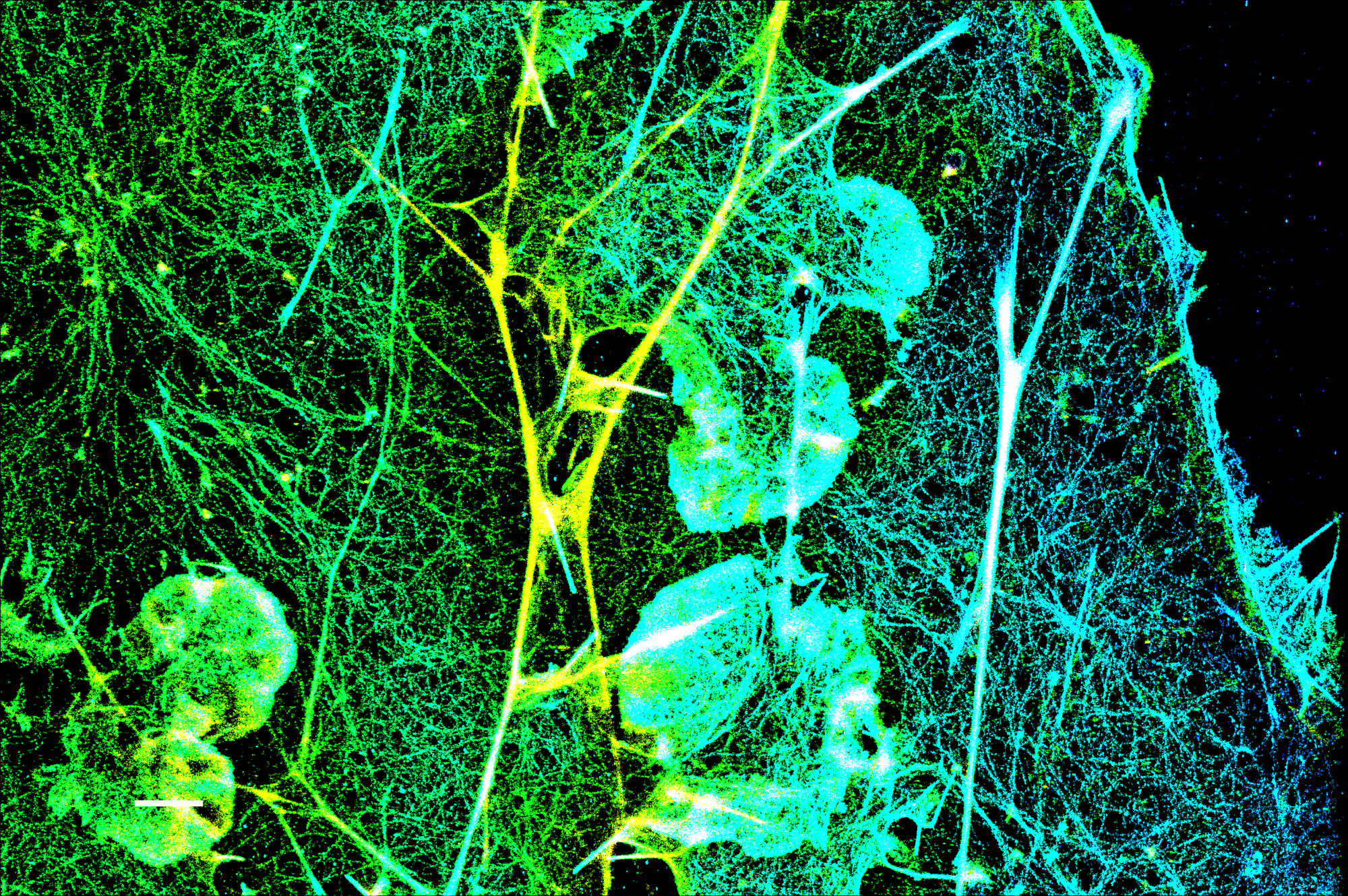
Structured Illumination Visualizes Patterns and is Ideal for Live Imaging
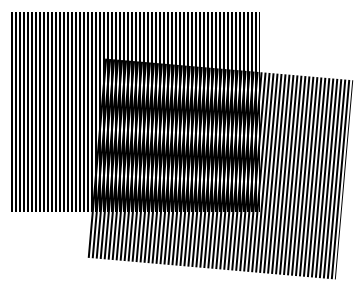
Structured illumination microscopy (SIM) uses a moveable grating to visualize patterns and structures beyond the diffraction limit. When two grids are misaligned, they create interference patterns known as Moiré fringes. If you know the identity of one grid, then you can mathematically deduce and reconstruct the second grid from the Moiré fringes. SIM uses excitation light focused into a striped pattern to make a grid. The sample acts as the unknown grid that is reconstructed.

SIM is ideal for super resolution imaging of live samples due to its fast acquisition and low illumination intensity, although its resolution is more modest than other super resolution techniques. SIM achieves a lateral resolution of 100 nm and optical sectioning with an axial resolution of about 300 nm. Another advantage of SIM is that it does not require specialized dyes, allowing super resolution imaging across the color spectrum using standard dyes. It is therefore trivial for a user to take a sample prepared for confocal or widefield microscopy and perform SIM.
SIM does have some caveats. Because the sample is reconstructed, the final images are not quantitative, therefore colocalization analysis of SIM images should not be done using correlations of pixel intensity, but rather object-based methods, such as measuring the overlap of segmented objects (see Hong et al., (2017)). Careful consideration of the biology and the reconstruction parameters is essential, as out of focus (‘honeycomb’) pseudo-structures are common, especially with non-punctate signal such as cell-fills. Lastly, SIM does not perform well when imaging deep sections of tissue, as scattering distorts the illumination and emission patterns.
Suppressing Fluorescence for a Sharper Image with STED
Stimulated emission depletion (STED) microscopy works similarly to beam scanning confocal but shrinks the size of the fluorescent spot using stimulated emission, producing a lateral resolution of about 50 nm. As with imaging, focusing the excitation beam on a sample is diffraction limited. STED uses a donut shaped depletion beam (also referred to as the STED beam) that encircles the excitation beam to reduce the excited area that fluoresces.
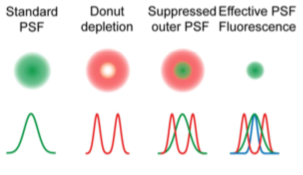
STED is useful for fixed and live imaging, but the imaging area must be reduced to have millisecond scale time resolution, and the high intensity STED beam can contribute to phototoxicity with prolonged imaging. Samples require minimal special preparation aside from the use of STED compatible fluorescent proteins or dyes like CF®405M, CF®488A, and CF®680R that can be depleted by the STED beam. Unlike other super resolution techniques, STED is quantitative because it does not rely on post-processing or reconstruction.
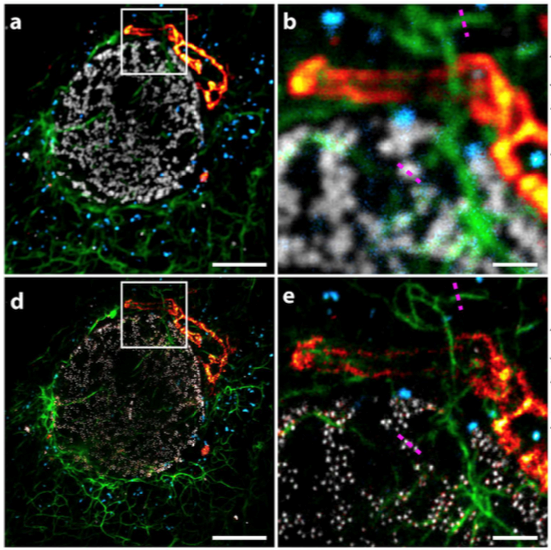
Improving STED with Superior Dyes and Lasers
STED is commonly limited to one or two STED channels per experiment because of the impracticality of using multiple STED beams. With advances in dye engineering, hyperspectral STED (“Hyper-STED”) has been developed to simultaneously image up to four dyes including CF®680R using a single STED beam (Winter et al., 2017). Coupled with spectral detection, this technique increases imaging speed and reduces phototoxicity. The white light supercontinuum laser used in Hyper-STED is currently cost-prohibitive, but accessibility should improve in the future with lower hardware costs and the availability of more STED compatible dyes.
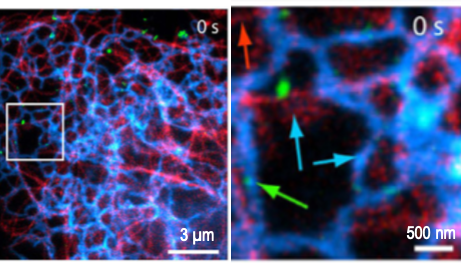
Imaging Single Molecules Gives Insight into Structure and Molecular Dynamics
The advent of bright and photostable dyes and fluorescent proteins has enabled a new era of single molecule imaging to uncover ultrastructure and the dynamic behavior of individual molecules. By collecting photons from isolated spots, single molecule techniques calculate the molecule’s most likely position within a resolution of 20 nm. Stochastic optical reconstruction microscopy (STORM) relies on random photoswitching of dyes between light and dark states, while its sibling, photoactivation localization microscopy (PALM), uses photoactivation of assemblies of isolated single molecules to construct an image. Single particle tracking (SPT) shows us the dynamic behavior of molecules as they move.
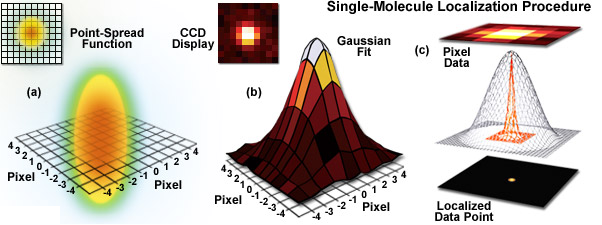
STORM Provides Unparalleled Resolution at a Cost
STORM offers the highest resolution in optical microscopy. By localizing single molecules, STORM constructs a composite image made of these localized spots by multiple cycles of photoactivation and bleaching. STORM has the intricate structure of intracellular assemblies, including the elegant axonal cytoskeleton, in a way that was not possible with conventional techniques. However, STORM suffers from low acquisition speed and phototoxicity due to the sample’s constant exposure to the intense photoactivation laser.
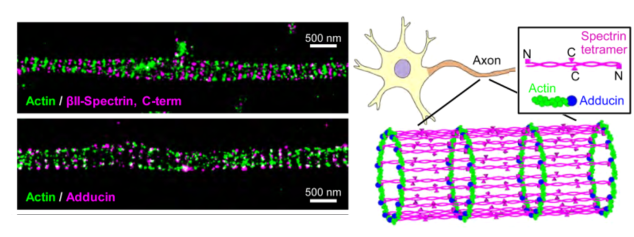
STORM uses photoswitching, “blinking” dyes, such as several CF® dyes, that have distinct light and dark states. While live imaging with STORM is possible, it is best applied to fixed samples where sufficient imaging cycles can be performed or live samples where object movement is minimal. Buffers, labeling, and mounting conditions need to be carefully optimized to promote dye switching. As with other super resolution techniques, dye innovations are driving improvements in STORM. STORM relies on intense and constant illumination, and multicolor STORM requires dyes that can be spectrally separated. Spectrally distinct, blinking dyes with improved molecular brightness reduce the intensity of light illumination and phototoxicity, and allow multicolor STORM with simple organic dyes.
Single Molecule Tracking Reveals Molecular Behavior
While STORM is used to gain structural insights, single molecule tracking (SMT) studies the dynamic behavior of molecules. Live imaging experiments using STED, SIM, or conventional microscopy image the average behavior of the population of labeled molecules, but SMT experiments allow the study of individual molecules, yielding valuable insights such as the existence of distinct populations of diffusing molecules and molecular partners.
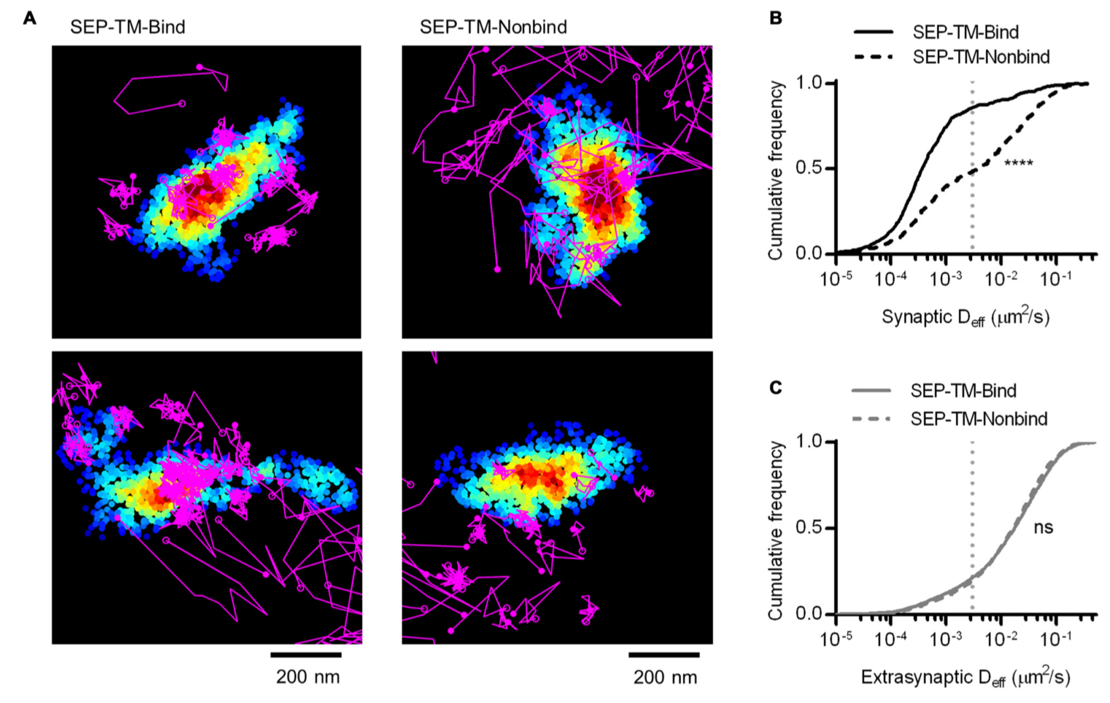
Single molecule tracking relies on photostable fluorophores. Quantum dots have been used due to their high brightness and photostability but are large compared to organic dyes and fluorescent proteins. Development is focused on producing photostable organic dyes and fluorescent proteins with high molecular brightness and low rates of photobleaching. The CF® dyes CF®633 and CF®680R are ideal for single molecule tracking, as their red and far-red excitation reduces phototoxicity.
Imaging at Depth with 2-Photon Microscopy
Super resolution and conventional microscopy suffer when attempting to image deep into tissue or behaving organisms. Visible light is absorbed or scattered by tissue, limiting its penetration, but infrared and near-IR light is less affected. While traditional single-photon microscopy excites a fluorophore with an absorbed wavelength of light, 2-photon microscopy instead uses two longer wavelength photons. 2-photon microscopy is commonly used at the cellular or macroscopic level with long working distance objectives to image organ-level organization with dyes including CF®488A, CF®594, and CF®680R.
2-photon microscopy is also commonly used to image biosensors, such as calcium and recently, ultrafast imaging of dopamine release (Patriarchi et al., 2018). Thus, developments in 2-photon microscopy center on acquisition speed and improving resolution at the cellular scale through dye engineering. While 2-photon microscopy can better penetrate tissue to excite dye molecules, emitted fluorescence is also subject to scattering. To that end, red-shifted dyes like CF®680R have been developed to improve resolution in deep imaging.
Super Resolution Trends Going Forward
Super resolution microscopy continues to improve. Key areas for improvement are in dyes and sample preparation. Recently, SIM has been combined with STORM principles to extend its resolving ability further (D. Li et al., 2015). Super resolution microscopy has been combined with new sample preparation methods like expansion microscopy to look even deeper into molecular organization (Tillberg et al., 2016; Wang et al., 2018; H. Xu et al., 2019).
References
- Dodgson, J., Chessel, A., Cox, S., & Carazo Salas, R. E. (2015). Super-Resolution Microscopy: SIM, STED and Localization Microscopy. In T. E. S. Dahms & K. J. Czymmek (Eds.), Advanced Microscopy in Mycology (pp. 47–60). Springer International Publishing. https://doi.org/10.1007/978-3-319-22437-4_3
- Hiersemenzel, K., Brown, E. R., & Duncan, R. R. (2013). Imaging Large Cohorts of Single Ion Channels and Their Activity. Frontiers in Endocrinology, 4. https://doi.org/10.3389/fendo.2013.00114
- Hong, S., Wilton, D. K., Stevens, B., & Richardson, D. S. (2017). Structured Illumination Microscopy for the Investigation of Synaptic Structure and Function. Methods in Molecular Biology (Clifton, N.J.), 1538, 155–167. https://doi.org/10.1007/978-1-4939-6688-2_12
- Gustafsson, M. G. L., Betzig, E., Hess, H. F., Patterson, G. H., Lippincott-Schwartz, J., & Davidson, M. W. (n.d.). ZEISS Microscopy Online Campus | Introduction to Superresolution Microscopy. Zeiss Education in Microscopy and Digital Imaging. Retrieved July 7, 2020, from http://zeiss-campus.magnet.fsu.edu/articles/superresolution/introduction.html
- Li, D., Shao, L., Chen, B.-C., Zhang, X., Zhang, M., Moses, B., Milkie, D. E., Beach, J. R., Hammer, J. A., Pasham, M., Kirchhausen, T., Baird, M. A., Davidson, M. W., Xu, P., & Betzig, E. (2015). ADVANCED IMAGING. Extended-resolution structured illumination imaging of endocytic and cytoskeletal dynamics. Science (New York, N.Y.), 349(6251), aab3500. https://doi.org/10.1126/science.aab3500
- Li, T. P., & Blanpied, T. A. (2016). Control of Transmembrane Protein Diffusion within the Postsynaptic Density Assessed by Simultaneous Single-Molecule Tracking and Localization Microscopy. Frontiers in Synaptic Neuroscience, 19. https://doi.org/10.3389/fnsyn.2016.00019
- Patriarchi, T., Cho, J. R., Merten, K., Howe, M. W., Marley, A., Xiong, W.-H., Folk, R. W., Broussard, G. J., Liang, R., Jang, M. J., Zhong, H., Dombeck, D., Zastrow, M. von, Nimmerjahn, A., Gradinaru, V., Williams, J. T., & Tian, L. (2018). Ultrafast neuronal imaging of dopamine dynamics with designed genetically encoded sensors. Science, 360(6396), eaat4422. https://doi.org/10.1126/science.aat4422
- Tillberg, P. W., Chen, F., Piatkevich, K. D., Zhao, Y., Yu, C.-C. (Jay), English, B. P., Gao, L., Martorell, A., Suk, H.-J., Yoshida, F., DeGennaro, E. M., Roossien, D. H., Gong, G., Seneviratne, U., Tannenbaum, S. R., Desimone, R., Cai, D., & Boyden, E. S. (2016). Protein-retention expansion microscopy of cells and tissues labeled using standard fluorescent proteins and antibodies. Nature Biotechnology, 34(9), 987–992. https://doi.org/10.1038/nbt.3625
- Wang, Y., Yu, Z., Cahoon, C. K., Parmely, T., Thomas, N., Unruh, J. R., Slaughter, B. D., & Hawley, R. S. (2018). Combined expansion microscopy with structured illumination microscopy for analyzing protein complexes. Nature Protocols. https://doi.org/10.1038/s41596-018-0023-8
- Winter, F. R., Loidolt, M., Westphal, V., Butkevich, A. N., Gregor, C., Sahl, S. J., & Hell, S. W. (2017). Multicolour nanoscopy of fixed and living cells with a single STED beam and hyperspectral detection. Scientific Reports, 7(1), 46492. https://doi.org/10.1038/srep46492
- Xu, H., Tong, Z., Ye, Q., Sun, T., Hong, Z., Zhang, L., Bortnick, A., Cho, S., Beuzer, P., Axelrod, J., Hu, Q., Wang, M., Evans, S. M., Murre, C., Lu, L.-F., Sun, S., Corbett, K. D., & Cang, H. (2019). Molecular organization of mammalian meiotic chromosome axis revealed by expansion STORM microscopy. Proceedings of the National Academy of Sciences, 116(37), 18423–18428. https://doi.org/10.1073/pnas.1902440116
- Xu, K., Zhong, G., & Zhuang, X. (2013). Actin, Spectrin, and Associated Proteins Form a Periodic Cytoskeletal Structure in Axons. Science, 339(6118), 452–456. https://doi.org/10.1126/science.1232251