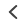Fluorescence microscopy is an indispensable tool for studying complex biological systems. However, the ability to distinguish higher numbers of discrete subcellular structures concurrently is hindered by the intrinsically broad spectra of fluorophores. Spectral imaging and unmixing, which entails resolving the measured spectrum of a mixed pixel into its constituent spectra and corresponding proportions, extends the possibilities to discriminate distinct fluorophores with overlapping emission spectra and, therefore, the capabilities of multi-color imaging. The number of channels available for analysis limits the number of fluorophores that can be unmixed, however using a combination of excitation and emission unmixing increases the number of fluorophores that can be analyzed in parallel. While several spectral imaging pipelines are emerging for expanded multiplex unmixing, they are often time-consuming, inefficiently utilize signal, or require expensive specialized equipment.
In a Biomedical Optics Express article from November 2022, Chen, Li, and Xu extend their previously published excitation spectral microscopy approach to the concurrent imaging of up to ten targets in simulations and eight in cell-imaging experiments. Unlike in conventional unmixing, where each fluorophore is identified by its emission spectra at fixed excitation wavelength, in this technique a fixed emission wavelength is used to collect the excitation spectrum for each fluorophore using a variable excitation source. The authors established a 24-wavelength excitation scheme through synchronized scanning of the excitation wavelength from a white lamp via an acousto-optic tunable filter (AOTF) so that full-frame excitation-spectral images are obtained every 24 frames for multiplexing a greater number of fluorophores or probes (Figure 1). AOTFs rely on a specialized birefringent crystal whose optical properties vary with changes in acoustic frequency, enabling very rapid wavelength tuning and simultaneous modulation of intensity and wavelength. This approach bypasses the limitations on multiplexing imposed by the number of detection channels, and enables faster live cell imaging by eliminating the time needed for filter switching. Using computational simulations of their system, the team established the feasibility of concurrent low-crosstalk imaging for ten commercial fluorophores with substantial spectral overlap spanning the visible range. Similarly, for cell-imaging experiments, they labeled different subcellular structures with eight of the ten fluorophores from the simulation and demonstrated the resolution of all eight structures with minimal color crosstalk. These labels included four Biotium dyes: LipidSpot™ 488, CF®514, CF®583R, and CF®620R. The authors thus showcase a relatively simple and cost-effective system for visualizing subcellular components with more colors and low crosstalk.
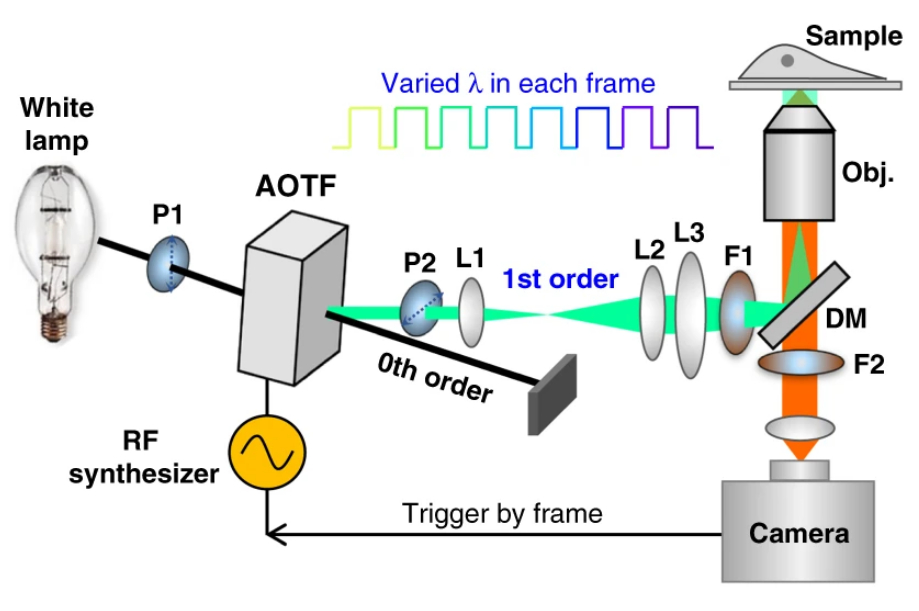
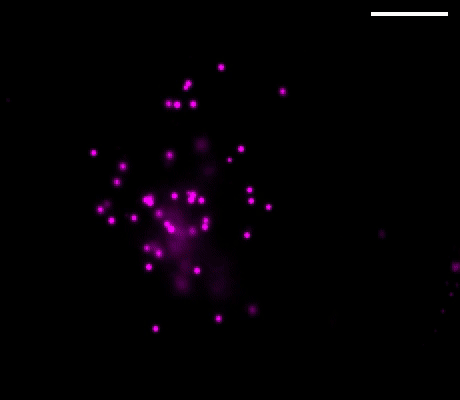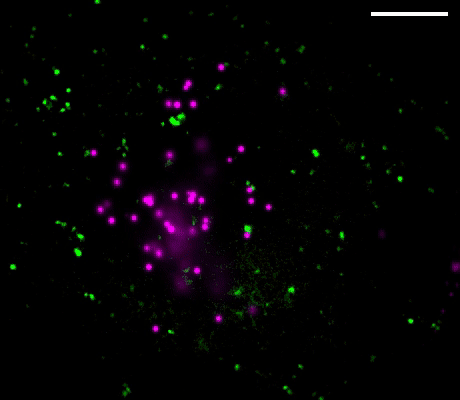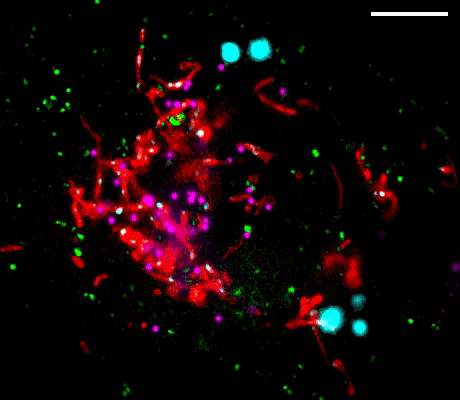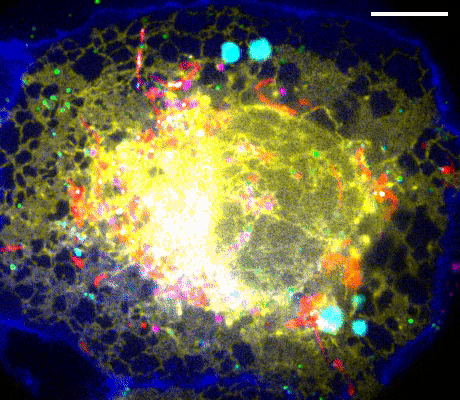
Figure 1. Left: Overview of the setup. Full-frame spectral micrographs are obtained by modulation of the excitation wavelength λ in consecutive frames. P polarizer, L lens, F bandpass filter, DM dichroic mirror. The resulting excitation spectrum collected at each pixel is linearly unmixed into the abundances of different fluorophores based on their pre-calibrated excitation spectrum. Right: Overlay of six fluorescently stained subcellular targets, including cell membrane stained using Biotium’s WGA-CF®532 and lipid droplets stained with Biotium LipidSpot™ 488, in a live COS-7 cell via 8-excitation-wavelength recording (From a 2021 publication by the same research group). Scale bar: 10 µm Credit: Modified from Chen, Yan, Xiang & Xu (2021) https://doi.org/10.1038/s41377-021-00536-3 reproduced under the Creative Commons license.
Learn more about Biotium’s high-performance CF® Dyes with multi-color flexibility, Mix-n-Stain™ Antibody Labeling Kits, and reagents for immunofluorescence microscopy. We also sell a wide variety of nuclear, organelle, and cell membrane stains.
Full Citation
Chen, K., Li, W., & Xu, K. (2022). Super-multiplexing excitation spectral microscopy with multiple fluorescence bands. Biomedical Optics Express, 13(11), 6048-6060. https://doi.org/10.1364/BOE.473241