EvaRuby™ Dye, 20X in Water
Visible red fluorescent DNA-binding dye for qPCR, HRM®, and LAMP. Can be combined with probe-based qPCR reactions with little cross-talk in other channels.
Please fill in the inquiry form and we will contact you shortly.
Wishlist updated! View wishlist
Product Description
EvaRuby™ is the first visible red DNA-binding dye for both qPCR and HRM. The dye has unique spectral properties that minimize cross-talk between channels. EvaRuby™ can be used as a typical intercalating dye for qPCR, HRM®, and LAMP. The dye also can be incorporated into probe-based qPCR assays to monitor the reaction in real-time and perform post-qPCR high-resolution melt analysis.
- Visible red DNA-intercalating dye for qPCR
- Combine probe-based qPCR with HRM® in the same reaction
- Unique spectral properties, for minimal cross-talk in other channels
- Use as an internal control to monitor probe-based qPCR reactions in real-time
- Avoid wasting time and resources on post-qPCR gel electrophoresis
- Ex/Em 480/613 nm (with DNA); detect with ~488 nm excitation and ~610 nm emission filters
The unique properties of EvaRuby™ Dye
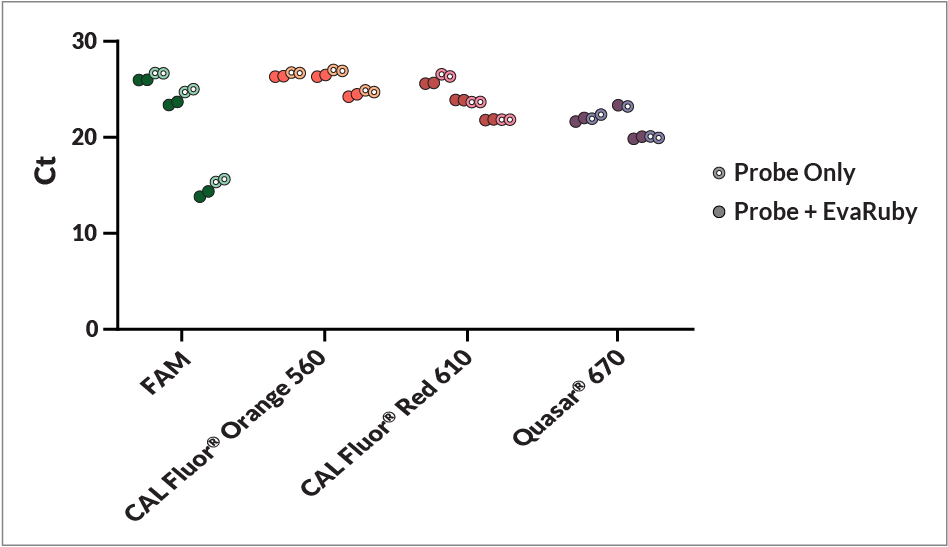
EvaRuby™ is a visible red fluorescent nucleic acid dye designed for qPCR, as such it is essentially non-fluorescent by itself, but becomes fluorescent upon binding to dsDNA. Unlike EvaGreen® or SYBR® intercalating qPCR dyes, EvaRuby™ has a long Stokes shift and orange fluorescent emission. With excitation/emission peaks at 480/613 nm, EvaRuby™ Dye is excited using the same green/blue channel as EvaGreen® but is detected in the orange (JUN™ or ROX) channel. As a result, EvaRuby™ has minimal cross-talk in other detection channels and therefore can be incorporated into singleplex and multiplex probe-based qPCR assays (Figure 1).
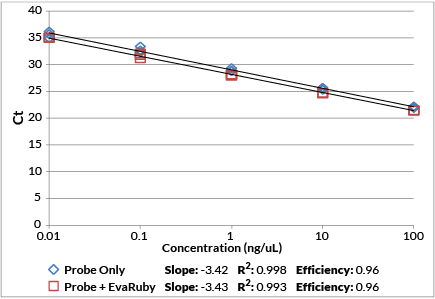
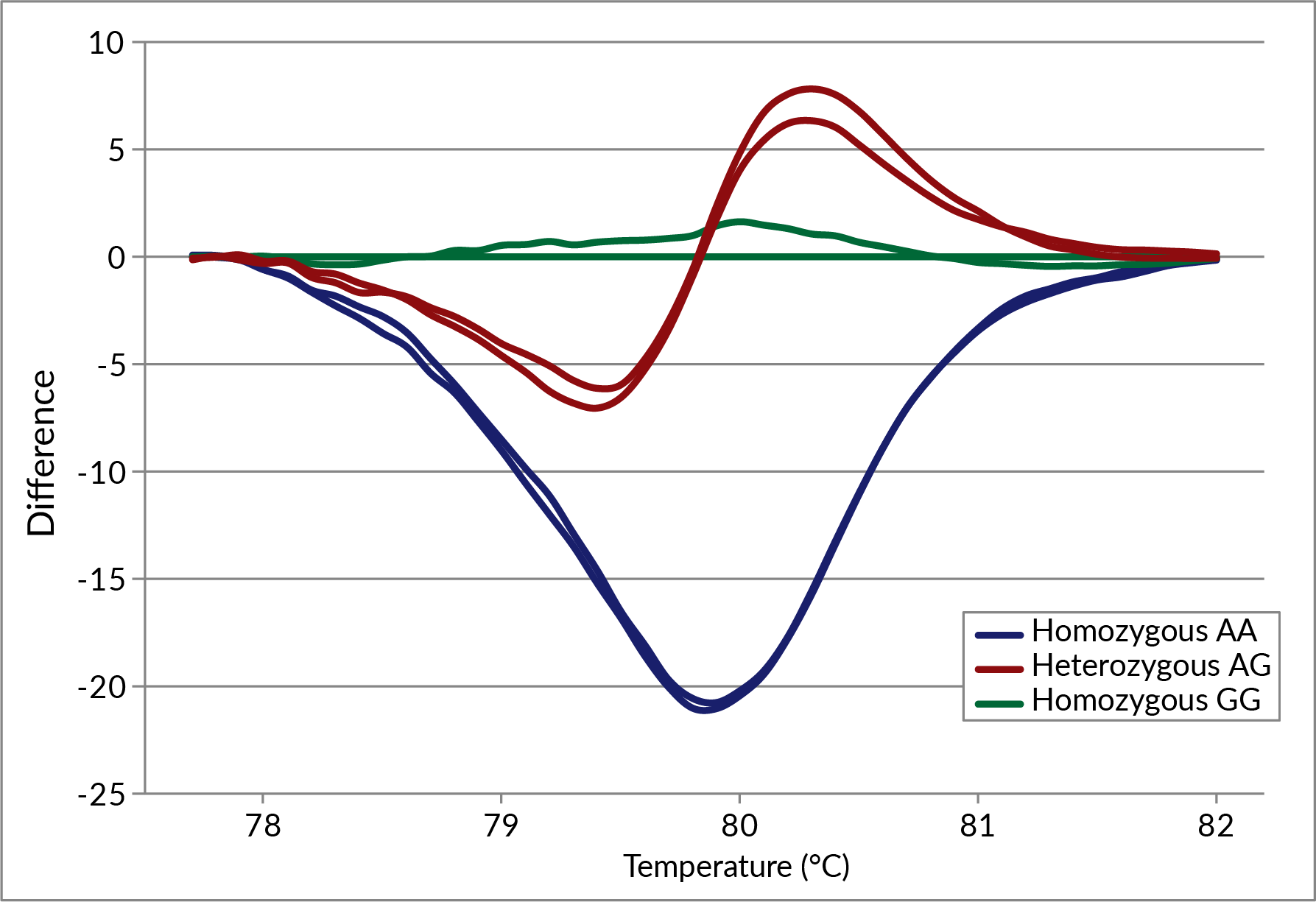
While EvaRuby™ can be used as a traditional intercalating qPCR dye to perform real-time qPCR, melt curve analysis, LAMP, and HRM®, the dye’s main advantage is that it can be incorporated into probe-based qPCR assays. Unlike other intercalating dyes, EvaRuby’s™ unique spectral properties allow it to be combined with commonly used fluorescent qPCR probes with little cross-talk in other fluorescence channels. The dye can be added to the reaction mix during reaction setup without affecting probe specificity (Figures 1, 2) while expanding the capabilities of probe-based qPCR to include real-time troubleshooting and post-qPCR high-resolution melt analysis (Figure 3). Note that for traditional single-color dye-based qPCR, we still recommend using EvaGreen® Dye.
A New Internal Control for Probe-based qPCR
Dye-based qPCR has low target specificity but has the advantage of performing a rapid and facile confirmation of results with melt curve analysis. On the other hand, probe-based qPCR benefits from high target specificity but is not compatible with melt curve analysis. Instead, confirmation of the qPCR results is often done by gel electrophoresis, a labor-intensive step that increases the time to results. Using EvaRuby™ in combination with probe-based qPCR enables the use of probe for high target specificity together with an intercalating dye for melt analysis and confirmation of results. Furthermore, because EvaRuby™ signal is independent from probe signal, it acts as an internal control for PCR amplification. For example, despite successful amplification, a degraded or suboptimal probe may fail to bind to the amplified target sequence and as a result, qPCR would incorrectly report amplification failure. However, with EvaRuby™ present, amplification would still be detected during qPCR and can be identified with melt curve analysis.
Instrument Compatibility
EvaRuby™ requires detection using a custom qPCR detection channel. Your qPCR instrument must have ~470 nm excitation and ~610 nm emission filters, as well as the option to acquire data using this non-standard filter combination. Instruments with paired excitation/emission filters may not be compatible with the dye. EvaRuby™ has been validated on Applied Biosystems® QuantStudio™ 5 96-well and Qiagen® Rotor-Gene® Q instruments. A compatibility table for other instruments is available in the PI; however it is important to confirm compatibility by checking your instrument technical specifications or by contacting the instrument manufacturer before purchasing EvaRuby™ Dye.
If your qPCR instrument requires a passive reference dye, use VeriFluor™ Far-Red (Cat. no. 29087) for optimal results. VeriFluor™ Far-Red is detected in the red (Cy®5 or Mustang Purple™) fluorescence channel. High ROX concentrations (~500 nM) are incompatible with EvaRuby™. Using low ROX concentrations (~50 nM) for normalization is suboptimal and will result in a small increase of the background fluorescence in the EvaRuby™ channel.