The International Space Station (ISS) is a closed system that contains diverse microorganisms, some that have been there from the beginning 17 years ago, and others introduced periodically via the crew and cargo. Since the ISS inception, microbes in the ISS have been monitored using culture-based methods. In a recent issue of Microbiome, Sielaff et. al. from the Jet Propulsion Laboratory in Pasadena report their extensive genomic analysis of bacterial and fungal populations in the ISS.
The researchers collected samples during three flight missions, in eight locations over a time span of 14 months to look for population changes over time and location. They performed strain identification using the current method of plate-based culturing, and compared that with the more sensitive method of qPCR and amplicon sequencing. Targeting the 16S gene for bacteria and the ITS region for fungi, they performed viability PCR using Propidium Monoazide (PMA), followed by sequencing and taxonomic analysis.
The total bacteria load did not change much between flights, or from site to site, although the total fungal load decreased over time. No differences between the PMA-treated and –untreated profiles were found, suggesting that most of the samples collected were viable organisms. Some of the identified species were pathogenic bacteria and fungi. The bacteria data sets were all dominated by the Enterobacteriaceae family, and the population profiles resembled the profiles of other human-populated environments. Interestingly, changes were observed over time, but not over location: the profile of different taxa was different for each of the three flights, but not statistically different between the eight locations.
This study lays the groundwork for subsequent long-term monitoring of the ISS microbiome, as well as potentially other unique environments.
Checinska Sielaff, A.,Urbaniak, C.,Mohan, G. B. M.,Stepanov, V. G.,Tran, Q.,Wood, J. M.,Minich, J.,McDonald, D.,Mayer, T.,Knight, R., et al. (2019). Characterization of the total and viable bacterial and fungal communities associated with the International Space Station surfaces. Microbiome 7, 50. https://doi.org/10.1186/s40168-019-0666-x
Learn more about viability PCR using PMA or PMAxx™.
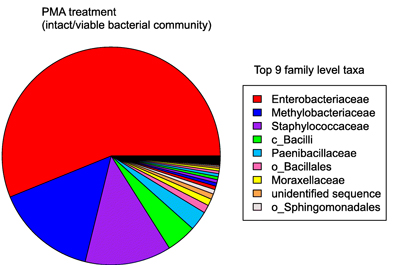
Note: Chart adapted from Characterization of the total and viable bacterial and fungal communities associated with the International Space Station surfaces. Checinska Sielaff, A., Urbaniak, C., et. al, 2019. Microbiome 7, 50. https://doi.org/10.1186/s40168-019-0666-x