Health benefits observed from the ingestion of bacteria in fermented food has fueled the use of probiotics in medicine and as health supplements. The survival and viability of ingested bacteria through the digestive system are key characteristics of effective probiotics. Lactobacilli species are resident microbes of the GI tract and widely used in food fermentation and probiotic preparations. Viability is traditionally estimated through time-consuming, low-throughput culture-based methods that poorly discriminate between closely-related bacterial species when present in the same sample. Quantifying viable but nonculturable bacteria is also challenging.
Molecular approaches like real-time qPCR can provide specific and accurate bacterial quantification. Droplet digital PCR (ddPCR) has emerged as an advancement over conventional qPCR for absolute, standard-free DNA quantitation and rare nucleic acid sequence detection. However, these approaches are non-discriminating between viable and non-viable bacteria. A recent paper in the Journal of Microbial Methods combines the sensitivity, specificity and speed of ddPCR with quantifiable viability by using the viability dye propidium monoazide (PMA) to measure viability of 3 Lactobacillus strains in piglet feces. All 3 lactobacilli strains were able to survive in the piglet GI tract. The authors were able to accurately and specifically quantify a small numbers of viable bacterial cells in the presence of a high-background of other microorganisms. Optimal conditions for PMA treatment and sample preparation to obtain accurate determination of viability are described. PMA efficiency differed even between the closely related species illustrating that PMA protocols need to be specifically adapted for each target strain. The PMA-ddPCR approach thus provides a new tool to quantify bacterial survival in fecal samples from preclinical and clinical samples.
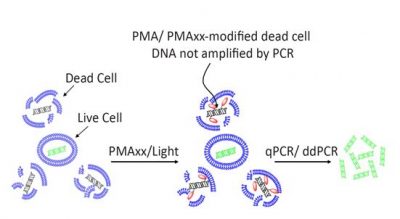
Principle of v-PCR with PMA/ PMAxx dyes.The membrane impermeable dyes selectively enter dead cells and modify their DNA. Modified DNA is not amplified, allowing quantitation of viable cell DNA.
To read the original article, click here.
Droplet digital PCR improves absolute quantification of viable lactic acid bacteria in faecal samples. Gobert G, Cotillard A, Fourmestraux C, Pruvost L, Miguet J, Boyer M. J Microbiol Methods. (2018) 148:64-73. doi:10.1016/j.mimet.2018.03.004.
Learn more about Biotium’s viability PCR (v-PCR) technology, v-PCR dyes PMA and PMAxx, and related v-PCR kits and reagents.