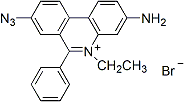
Ethidium Monoazide Bromide (EMA)
A photo-activated fluorescent nucleic acid binding dye used to selectively and covalently label dead cells in the presence of live cells for viability PCR, fluorescent microscopy, and flow cytometry.
Please fill in the inquiry form and we will contact you shortly.
Wishlist updated! View wishlist
Product Description
When not activated or bound to DNA, Ethidium monoazide bromide (EMA) is a non-fluorescent nucleic acid stain with a photoaffinity label. Once photoactivated, the dye binds covalently to nucleic acids and can emit fluorescence. The dye is most often used to label dead cells in a mixed population of live and dead cells.
- Covalently label DNA with a photo-activated fluorescent stain
- Cell membrane-impermeable
- Label dead cells in a mixed population of live and dead cells
- Compatible with fluorescent microscopy, fluorescence plate reader, or flow cytometry
- Orange solid soluble in DMF, or ethanol
Because EMA is relatively impermeant to live cells, it selectively binds DNA in dead cells. Photolysis renders the dead cell DNA covalently labeled with the dye. One can then wash and fix the cell preparation for analysis by microscopy, fluorescence plate reader or flow cytometry. The dye has been used to footprint drug binding sites on DNA, to modify plasmid DNA, and for labeling yeast cells for leukocyte phagocytosis assays. The major advantage of this method is that researchers can avoid extensive manipulation of live pathogenic organisms. EMA has been used to differentiate between viable and dead bacteria by 5′-nuclease PCR.
Viability PCR
Switching from agar plates to v-PCR is a great way to speed up bacteria viability testing. There are some things that you will need to know and take into consideration before you switch to the v-PCR method. For viability PCR applications, Ethidium monoazide (EMA) is the most cell-permeable, and may result in false negatives. Propidium monoazide (PMA), developed at Biotium, is the most widely-used dye, so there are many protocols and publications available for reference. PMAxx™ is designed be superior to PMA and EMA as well as compatible with existing PMA protocols. PMAxx™ has better differentiation of live and dead bacteria and therefore it is more selective for dead cell labeling for v-PCR quantitation of bacteria. See the PMA & PMAxx™ Reference List and PMA & PMAxx™ Validated Bacterial Strains. For more information about v-PCR, see our blog post.
Viability PCR Kits
To more easily get started with viability PCR, learn about our customizable Viability PCR Starter Kits. You choose which viability PCR dye you prefer, and whether or not you need Enhancer for gram-negative bacteria. All of the starter kits come with an EvaGreen® qPCR master mix and a detail protocol to get you started.
Biotium also provides strain-specific Real-Time PCR Bacterial Viability Kits with validated primers for selected pathogens: M. tuberculosis, S. aureus, MRSA, Salmonella, E. coli, E. coli O157:H7, Legionella and Listeria. These kits provide everything that you need for the selective detection of your favorite species of live bacteria by real-time v-PCR.
LED Photolysis Devices
Biotium sells two devices for the photolysis step of viability PCR. The PMA-Lite™ LED photolysis device is designed for PMAxx™-, PMA-, and EMA-treated samples in microcentrifuge tubes. The Glo-Plate™ Blue is a flat LED box ideal for samples in microplates, filters, or other formats.
 PMA-Lite™ LED photolysis device, for photoactivation of PMAxx™, PMA, or EMA
PMA-Lite™ LED photolysis device, for photoactivation of PMAxx™, PMA, or EMA
 Glo-Plate™ Blue LED Illuminator for photoactivation of PMAxx™, PMA, and EMA
Glo-Plate™ Blue LED Illuminator for photoactivation of PMAxx™, PMA, and EMA
References
- Plant Dis, 104, 1105, (2020), DOI: 10.1094/PDIS-10-19-2248-RE
- Environ. Sci.: Water Res. Technol., 6, 87, (2020), DOI: 10.1039/C9EW00896A
- J Leukoc Biol., Pre-Print, (2020), DOI: 10.1002/JLB.5MA1219-296R
- Cell, 178, 1231, (2019), DOI: 10.1016/j.cell.2019.07.033
- Curr Microbiol, 76, 1425, (2019), DOI: 10.1007/s00284-019-01772-y
- Environ. Sci.: Water Res. Technol., 5, 91, (2019), DOI: 10.1039/C8EW00613J
- AIMS Microbiol., 5, 102, (2019), DOI: 10.3934/microbiol.2019.1.102
- Water, 11, 1196, (2019), DOI: 10.3390/w11061196
- J Microbiol Methods, 148, 33, (2018), DOI: 10.1016/j.mimet.2018.03.018
- Water, 10, 7, (2018), DOI: 10.3390/w10070841
- Food control, 85, 23, (2018), DOI: 10.1016/j.foodcont.2017.09.013
- Vaccine, 35, 4330, (2017), DOI: 10.1016/j.vaccine.2017.06.082
- Breast Cancer Res., 19, 20, (2017), DOI: 10.1186/s13058-017-0813-x
- Appl Microbiol Biotechnol., 101, 7371, (2017), DOI: 10.1007/s00253-017-8471-6
- J Control Release., 265, 66, (2017), DOI: 10.1016/j.jconrel.2017.04.027
- J Mol Biol. (1975), DOI: 10.1016/0022-2836(75)90231-4