A Powerful PCR-Based Method for Analyzing Microbial Viability
Introduction to Viability PCR (v-PCR)
Reliable detection of living microbes and viruses is of widespread interest across numerous sectors including basic research, food safety, agriculture, and public health. However, traditional methods of measuring microbial viability rely on time-consuming culturing, colony counting, or microscopic observation. In addition, certain organisms cannot be cultured with current technology. Consequently, these viable but non-culturable (VBNC) species require molecular measures of viability. Viability PCR (v-PCR) is a reliable and culture-free method that offers sensitive and rapid detection of viable microorganisms. Since 2006, Biotium has pioneered the field of v-PCR with the development of PMA (Propidium Monoazide) and the improved PMAxx™ v-PCR dye. These photoreactive, membrane-impermeant, DNA binding dyes offer superior dead cell selectivity over traditional culture-based methods and the previous EMA (Ethidium Monoazide) v-PCR dye.
Advantages of v-PCR with PMA & PMAxx™
- Sensitive and rapid viability analysis without time-consuming culturing, microscopy, or colony counting
- Allows strain-specific viability analysis
- Validated and published for dozens of bacteria, fungi and viral species
- Compatible with Next-Generation Sequencing applications
- Suitable for complex sample types including soil, feces, water/wastewater, biological specimens, and food
An Industry Leader in v-PCR
PMA is the v-PCR dye of choice among industry leaders, and has been validated in hundreds of peer-reviewed publications and numerous strains of bacteria, fungi, and viruses. Early studies focused on v-PCR in bacteria, validating the procedure in dozens of different bacterial strains such as E. coli, Listeria, Legionella, Lactobacillus, M. tuberculosis, Chlamydia, H. pylori, and many other strains with relevance to human health. v-PCR has been most widely used in bacteria, but there are also dozens of publications using v-PCR in yeast and fungi, viruses, as well as a variety of other cell types such as archaea, amoeba, and parasites.
Published Applications for Viability PCR
- Microbiome Studies
- Environmental Testing
- Probiotic Research
- Clinical Testing
- Food & Water Sanitation
- Wine Fermentation
- Microbial Sequencing
Measuring Bacterial Viability in Environmental Samples?
Sign up to our e-newsletter to download our Literature Digest: Monitoring Viable Bacteria in the Environment with PMA & PMAxx™.
In this digest we have provided summaries and takeaways for three publications on the use of v-PCR with PMA and PMAxx™ for monitoring bacterial viability in samples taken from wastewater, fresh water, and drinking water treatment plants.
Looking to Study Viral Integrity with PMA or PMAxx™?
v-PCR is a proven method for monitoring capsid integrity for dozens of viral strains, including SARS-CoV-2.
The figure below displays the genus and strain number of viruses investigated for using v-PCR to monitor capsid integrity in peer-reviewed studies in the last decade (2009 to 2021) in the fields of food safety and environmental virology.
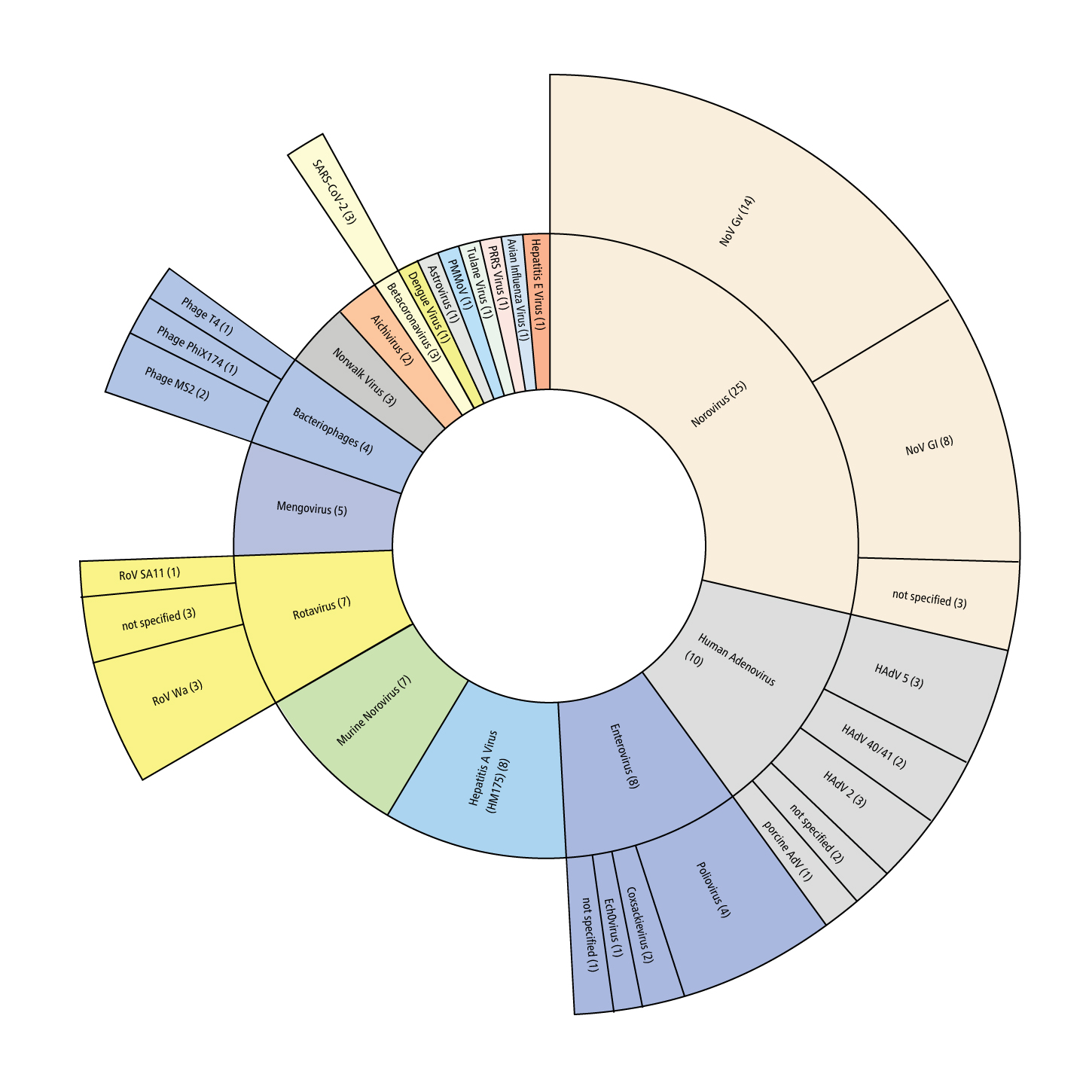
How v-PCR works
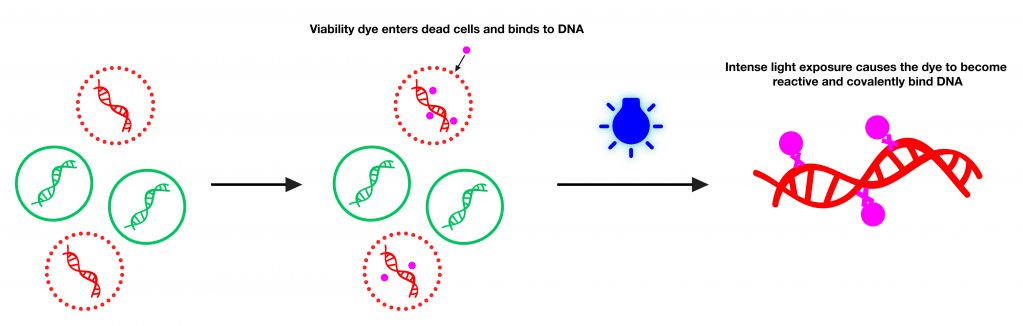
The infographic below shows the basic mechanism of v-PCR. PMAxx™ and PMA are photoreactive dyes with high affinity for DNA. The dyes intercalate into dsDNA and form a covalent linkage upon exposure to intense visible light. The reaction inhibits PCR amplification of modified DNA templates by a combination of removal of modified DNA during purification and inhibition of template amplification by DNA polymerases. Because the dyes are cell membrane-impermeable, when a sample containing both live and dead bacteria is treated with dye, only dead bacteria with compromised cell membranes are susceptible to DNA modification. In a qPCR reaction, dead cell DNA will show delayed amplification and higher Ct than live cells.
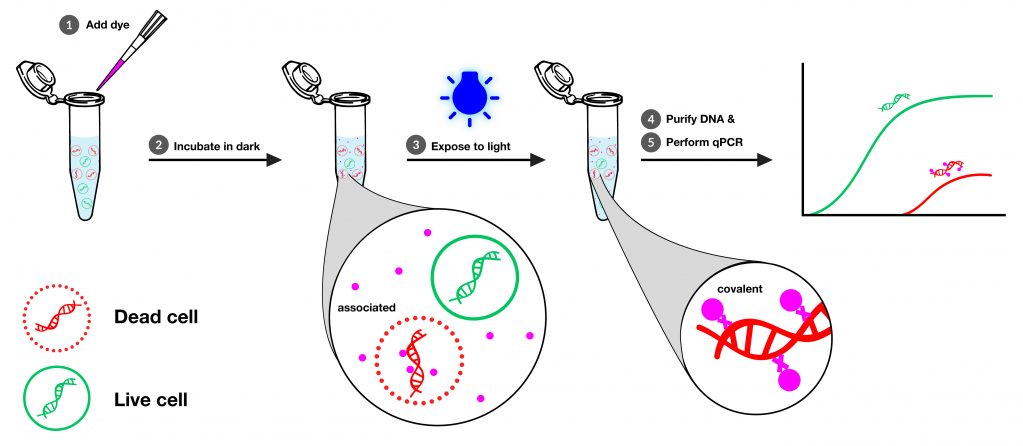
v-PCR Workflow
The basic workflow for v-PCR is shown in the infographic below (detailed protocols can be found on product pages). It is adaptable for many different sample types and experimental set-ups. For a small number of cultures, the samples can be prepared in microcentrifuge tubes, and exposed to light in the PMA-Lite™ 2.0 LED Photolysis Device (See lower on this page for more information).
Workflow Modifications
Many publications have used modifications of the basic protocol for specialty applications. For example, for large, dilute volumes, samples can be filtered onto a membrane, followed by dye treatment and photolysis on the membrane itself. Opaque, complex samples such as soil or feces generally need a higher dye concentration and longer light exposure; sample dilution can also help. And some users choose to perform qPCR directly on bacterial lysates, instead of purifying the DNA first.
PMA & PMAXX™: THE INDUSTRY STANDARD FOR VIABILITY PCR
PMA is validated in hundreds of journal articles for a wide variety of strains
Since Biotium first developed PMA, there have been hundreds of publications on the use of the dye in many sample types including dozens of bacterial strains, biofilms, yeast, fungi, viruses, and eukaryotic cells. It has been used in such applications as food and water safety and environmental testing, and has been used in conjunction with qPCR, NextGen Sequencing (NGS), Sanger sequencing, and Loop-mediated Isothermal Amplification (LAMP).
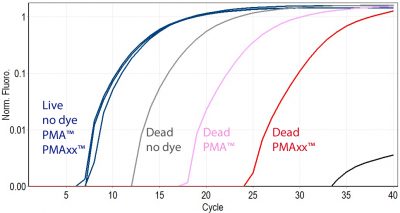
PMAxx™ is a superior alternative to PMA
While PMA is generally effective at differentiating between live and dead bacteria by qPCR, it does not completely eliminate PCR products from dead cell DNA. This could potentially give false positive results. PMAxx™ was designed by Biotium scientists to be a superior alternative to PMA. PMAxx™ is much more effective at eliminating PCR amplification of dead cell DNA, and therefore provides the best discrimination between live and dead bacteria. Learn more about viability PCR with PMA and PMAxx™ with links to our flyer and our full list of references below.
| Product Name | Catalog No. | Features |
|---|---|---|
| PMAxx™, 20 mM in H2O | 40069 | • Next-generation v-PCR dye • Gives the best live/dead discrimination |
| PMA Dye | 40013 | • Original v-PCR dye developed at Biotium • Validated in hundreds of publications and a wide variety of organisms |
| PMA Dye, 20 mM in H2O | 40019 |
STARTER KITS FOR V-PCR
The customizable Viability PCR Starter Kits contain the materials that you need for selective detection of viable cells using either PMA or PMAxx™ viability dye and qPCR. These kits can be used with any cell type of your choosing (see the PMA/PMAxx™ Reference List and Validated Bacterial Strains List for published validated cell types). All of the kits contain Forget-Me-Not™ EvaGreen® qPCR Master Mix, and your choice of PMA or PMAxx™ dye. You also have the choice of selecting a kit with or without the Enhancer for Gram Negative Bacteria (not to be used with other cell types). The user will need to supply their own primers to amplify their species of interest (see this blog post for primer design considerations). Each kit contains reagents sufficient to treat 80 bacterial cultures and perform 200 PCR reactions.
Kits include:
• PMA dye OR PMAxx™ dye
• Forget-Me-Not™ EvaGreen® qPCR Master Mix
• ROX reference dye
• PMA Enhancer (gram-negative strains only)
Not included but required:
• primers to amplify your cell type of interest
• DNA extraction kit or reagents
| Product Name | Catalog No. | Viability Dye | Target Cell Type |
|---|---|---|---|
| Viability PCR Starter Kit with PMA | 31075 | PMA | Any cells |
| Viability PCR Starter Kit with PMAxx™ | 31075-X | PMAxx™ | Any cells |
| Viability PCR Starter Kit with PMA and Enhancer | 31076 | PMA | Gram-negative bacteria |
| Viability PCR Starter Kit with PMAxx™ and Enhancer | 31076-X | PMAxx™ | Gram-negative bacteria |
PMA ENHANCER FOR GRAM-NEGATIVE BACTERIA
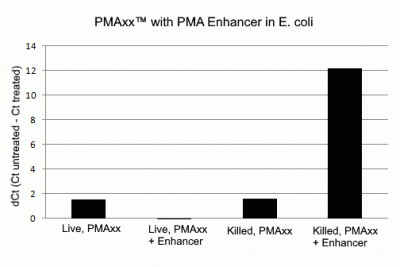
PMA Enhancer for Gram Negative Bacteria was designed to improve discrimination between live and dead gram-negative bacteria during viability PCR. PMA Enhancer is provided as a 5X solution, and is added to a sample before the addition of viability dye. When a sequence from a gram-negative bacteria is amplified by PCR, samples pre-treated with Enhancer show a decrease in the signal from dead cells, with no change in the signal from live cells. PMA Enhancer is compatible with PMAxx™ as well as PMA. PMAxx™ plus Enhancer is the optimal way to perform viability PCR on gram-negative bacteria. PMA Enhancer is available as a stand-alone product, and is also a component of our PMA Bacterial Viability Kits (gram-negative strains only) (see below).
The mechanism by which Enhancer improves live/dead discrimination has not been determined. It may improve passive permeability of dead cell walls or membranes to the dye, and/or improve access of the dye to the dead cell DNA.
PMA Enhancer for Gram-Negative Bacteria
| Product Name | Catalog No. | Features |
|---|---|---|
| PMA Enhancer for Gram Negative Bacteria, 5X Solution | 31038 | • Improves live/dead discrimination in gram-negative strains |
PHOTOACTIVATION DEVICES
The Only Devices Designed for Photoactivation of v-PCR Samples
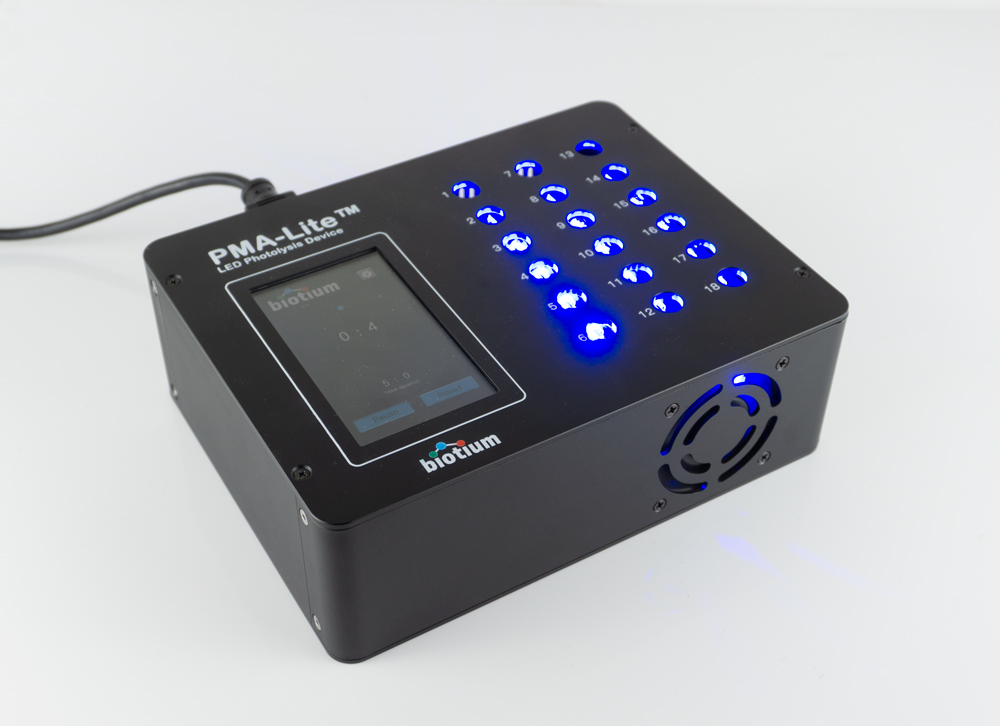
Biotium offers unique photoactivation devices designed for optimal photolysis of samples treated with PMAxx™ or PMA. The PMA-Lite™ 2.0 LED Photolysis Device may be used with microcentrifuge tubes, while the Glo-Plate™ 2.0 Blue LED Illuminator is designed for microplates or larger tubes. Both devices provide consistent and uniform illumination of samples in a temperature-controlled environment, for robust and reliable results.
For Microcentrifuge Tubes
PMA-Lite™ 2.0 LED Photolysis Device Features:
- Uniform illumination to up to 18 microcentrifuge tubes
- Efficient activation of v-PCR dyes such as PMAxx™ and PMA
- Timer for 5-45 minutes of photoactivation
- Long-lasting LED lights
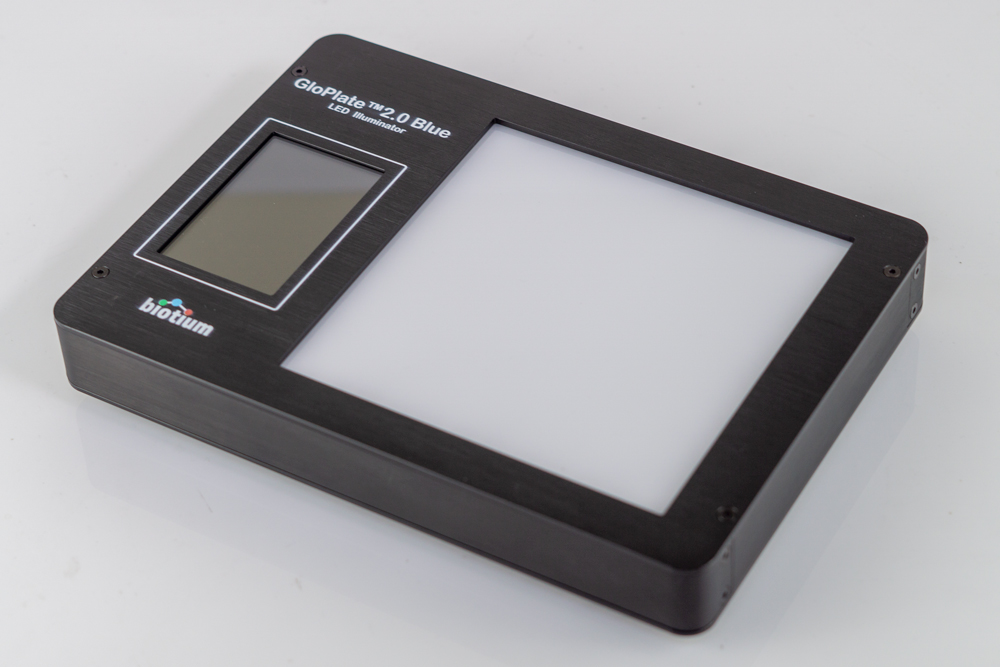
For Microplates and Large Tubes
Glo-Plate™ 2.0 Blue LED Illuminator Features:
- Uniform illumination for microplates, tubes, and gels
- Efficient activation of v-PCR dyes such as PMAxx™ and PMA
- Timer for 5-45 minutes of photoactivation
- Long-lasting LED lights
STRAIN-SPECIFIC V-PCR KITS
Strain-specific v-PCR kits are designed for selective detection of viable bacteria from a specific strain using PMAxx™ or PMA dye and real-time PCR. The kits contain PMA dye OR PMAxx™ dye, Forget-Me-Not™ EvaGreen® qPCR Master Mix, and validated PCR primers for detection of selected strains of bacteria that are of widespread interest to food safety, public health, and/or antibacterial research.
Don’t see your favorite bug? Let us know at techsupport@biotium.com
Kits include:
• PMA dye OR PMAxx™ dye
• Forget-Me-Not™ EvaGreen® qPCR Master Mix
• ROX reference dye
• PMA Enhancer (gram-negative strains only)
• Validated PCR primers for specific bacterial strain
PMA Real-Time PCR Bacterial Viability Kits are available for:
• Salmonella enterica
• Mycobacterium tuberculosis
• Staphylococcus aureus
• Staphylococcus aureus (methicillin-resistant)
• Listeria monocytogenes
• E. coli
• E. coli O157:H7
• Legionella pneumophila
Strain-Specific Bacterial Viability PCR Kits
| Bacteria Strain | Gene name | Kit with PMA (catalog #) | Kit with PMAxx™ (catalog #) |
|---|---|---|---|
| Salmonella enterica | invA | 31033 | 31033-X |
| Mycobacterium tuberculosis | groEL2 | 31034 | N/A |
| Staphylococcus aureus | nuc | 31035 | N/A |
| MRSA | mecA | 31036 | N/A |
| E. coli O157:H7 | Z3276 | 31037 | 31037-X |
| E. coli | uidA | 31050 | 31050-X |
| Listeria monocytogenes | hly | 31051 | 31051-X |
| Legionella pneumophila | mip | 31053 | N/A |
Associated products
PMA Enhancer for Gram Negative Bacteria, 5X Solution
PMA Real-Time PCR Bacterial Viability Kit – Listeria monocytogenes (hly)
31051, - 31051-XView allHide
PMAxx™ Dye, 20 mM in H2O
40069-1ML, - 40069View allHide
PMA Real-Time PCR Bacterial Viability Kit – E. coli O157:H7 (Z3276)
31037, - 31037-XView allHide




